A University of Toronto research team has created a new platform that delivers multiple therapeutic proteins to the body - each at its own independently controlled rate. The innovation could help treat degenerative diseases such as age-related macular degeneration (AMD), the leading cause of vision loss for people over 50.
Unlike traditional drugs made of small molecules, therapeutic proteins are synthetic versions of larger biomolecules naturally present in the body. One example is the synthetic insulin used to treat diabetes. There are other proteins that can modulate the body's own repair processes in ways that small-molecule drugs cannot.
"Proteins have great therapeutic potential, but they are notoriously difficult to deliver," says Molly Shoichet, a University Professor in the department of chemical engineering and applied chemistry in the Faculty of Applied Science & Engineering, the Institute for Biomedical Engineering and the Donnelly Centre for Cellular and Biomolecular Research, who led the team.
"For more than a decade, our group has been coming up with different ways to solve that fundamental problem."
One challenge is that proteins are less chemically stable than small molecules, making them vulnerable to damage from physical forces or interactions with solvents and other chemicals. Another challenge is getting them to the location where they are needed.
"You can inject a therapeutic protein to the site of an injury, but natural biochemical processes within the body tend to wick them away pretty quickly," says Carter Teal, a biomedical engineering PhD candidate and a graduate student in Shoichet's lab.
"To overcome that, you need systems that slowly release the proteins in a controlled way - and you can't necessarily use the same ones you would for traditional drug molecules, because they are often too harsh."
Shoichet and her team specialize in making biocompatible materials called hydrogels, which have a porous, sponge-like structure with spaces large enough to allow therapeutic proteins to move freely. Hydrogels can be infused with these proteins, then implanted into the body, where the proteins slowly diffuse out over time.
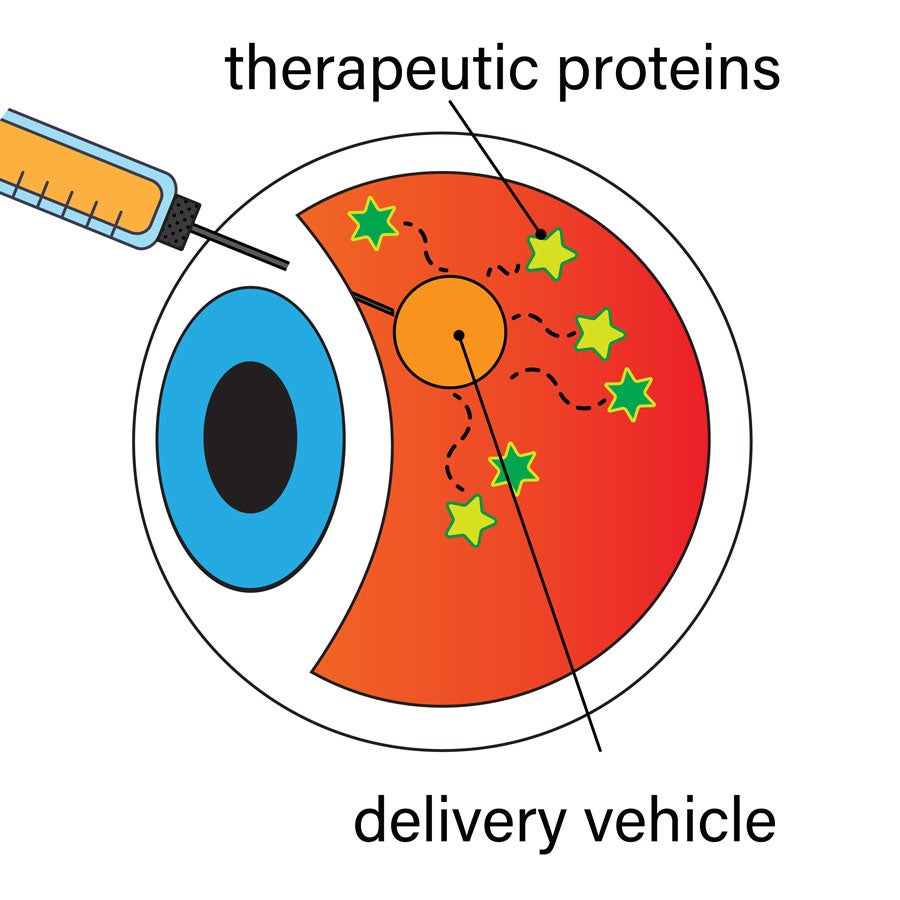 In this diagram, the orange circle represents a new biocompatible material that can deliver therapeutic proteins in a controlled manner to affected parts of the body such as the eye. The proteins, represented by green stars, can modulate natural biochemical processes that repair damage (illustration by Carter Teal)
In this diagram, the orange circle represents a new biocompatible material that can deliver therapeutic proteins in a controlled manner to affected parts of the body such as the eye. The proteins, represented by green stars, can modulate natural biochemical processes that repair damage (illustration by Carter Teal)
"One advantage of hydrogels is that they can be customized to provide desired mechanical properties and protein release rates through modification with a variety of chemical groups that cross-link in different ways or interact with the therapeutic proteins we want to deliver," says Marian Hettiaratchi, who worked on the project as a post-doctoral researcher in Shoichet's lab and is now a professor at the University of Oregon's Phil and Penny Knight Campus for Accelerating Scientific Impact.
"Proteins can also typically be easily mixed into hydrogels without disrupting their structure, and a number of hydrogels have been shown to protect proteins from degradation when delivered to the body."
In their latest paper, published in Advanced Materials, the team modified their hydrogels with affibody proteins, biomolecules that are akin to small antibodies.
"Affibody proteins were originally derived from a bacterium known as Staphylococcus aureus," says Teal.
"S. aureus produces a protein - known as protein A - and by separating and modifying a small section of protein A, customizable binding agents for many different kinds of biomolecules can be created. These binding agents are called affibody proteins."
The U of T Engineering team worked with University of Minnesota Twin Cities Professor Ben Hackel and his team in the department of chemical engineering and materials science, who provided a library of more than 100 million variant affibody proteins.
Shoichet's team then screened this library to find affibody proteins that would bind to each of the two therapeutic proteins they wanted to deliver: insulin-like growth factor-1 (IGF-1) and pigment epithelium-derived factor (PEDF).
Past research has shown that IGF-1 and PEDF each have the potential to slow the progress of retinal degenerative diseases like AMD on their own. When used in combination, they may be even more potent than either one by itself.
"Normally, when researchers do these types of screens, they are looking for affibody proteins with both high selectivity, meaning they bind only to the protein you want, and high affinity, meaning they bind tightly to the target protein," says Shoichet.
"In our case, we wanted high selectivity, but only moderate affinity, because we want the therapeutic protein to be slowly released over time. It's a bit like finding treasures in the trash: these are variants that, in a different context, would just have been thrown out."
After narrowing the field down to the two affibody proteins with the selectivity and affinity they needed, the team used a technique called "click chemistry" to coat them onto their hydrogels.
Tests conducted on these materials showed that IGF-1 and PEDF were released more slowly and stayed active in solution longer with these new affibody-hydrogels than with hydrogels without affibodies.
They also showed that the rates of release could be controlled independently by varying the levels of the two affibody proteins. For example, if the team wanted to slow the release rate of IGF-1 while leaving the release rate of PEDF unchanged, they could simply use more of the corresponding affibody protein pair in the hydrogel.
"The screening process has a lot of different steps in it, and it took many months to go from hundreds of millions of potential candidates down to the two we eventually selected," says Teal. "It was really exciting to see that, at the end of that, we achieved what we set out to do."
Because the approach is modular, Shoichet says it could easily be adapted for other uses.
"We chose IGF-1 and PEDF because of their synergistic effect in treating macular degeneration, which has long been a focus of our lab, but you could do this for any set of proteins," she says.
"Now that we've shown it's possible, we hope that it opens up the field for others."






