
As the world grapples with the challenge of containing the spread of the SARS-CoV-2 virus and healthcare professionals on the front lines battle the COVID-19 pandemic, scientists are hard at work studying the deadly virus to find solutions.
The KAUST Pathogen Genomics Laboratory (PGL), which is led by principal investigator Professor Arnab Pain, focuses on using high-throughput DNA and RNA sequencing-based methods and other functional genomic technologies to understand the biology and genomic diversity of pathogens that have a significant impact on human and animal health. Pain and his group are an integral part of the University's Rapid Research Response Team (R3T), which supports the Kingdom's healthcare stakeholders in combating the spread of COVID-19.
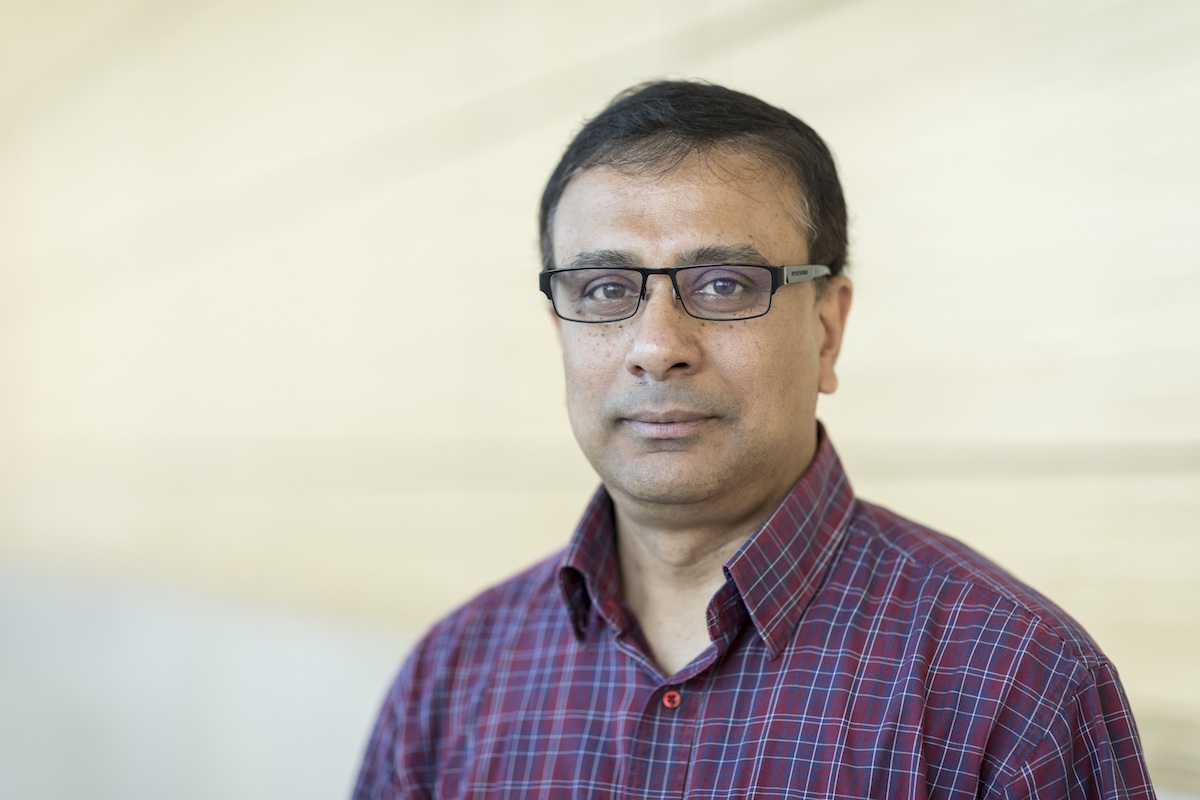

KAUST Professor Arnab Pain (pictured) and his Pathogen Genomics Laboratory are a critical piece of the University's efforts to combat the spread of COVID-19. Photo by Helmy Alsagaff.
Benefiting the Kingdom and the world
"As part of R3T, myself and my colleagues are trying to do as much as we can-using our expertise and the technologies we have access to-to come up with various potential solutions to help research and also provide some real assistance in the early detection and monitoring of this virus," Pain said.
Pain and Sharif Hala, one of his group's Ph.D. students, have been diligently setting up partnerships with the Saudi-CDC and several Ministry of Health (MoH) hospitals in Jeddah, Makkah and Madinah to facilitate the genome sequencing and bioinformatics analysis of a large collection of Saudi SARS-CoV-2 samples.
Hala has leveraged relationships around the Kingdom's medical community, including from his native Madinah, to establish a significant number of links with local hospitals and clinical doctors.
Apart from his role as a Ph.D. student, Hala is also an employee of King Abdullah International Medical Research Center, part of the Saudi National Guard Health Affairs-a tertiary medical complex.
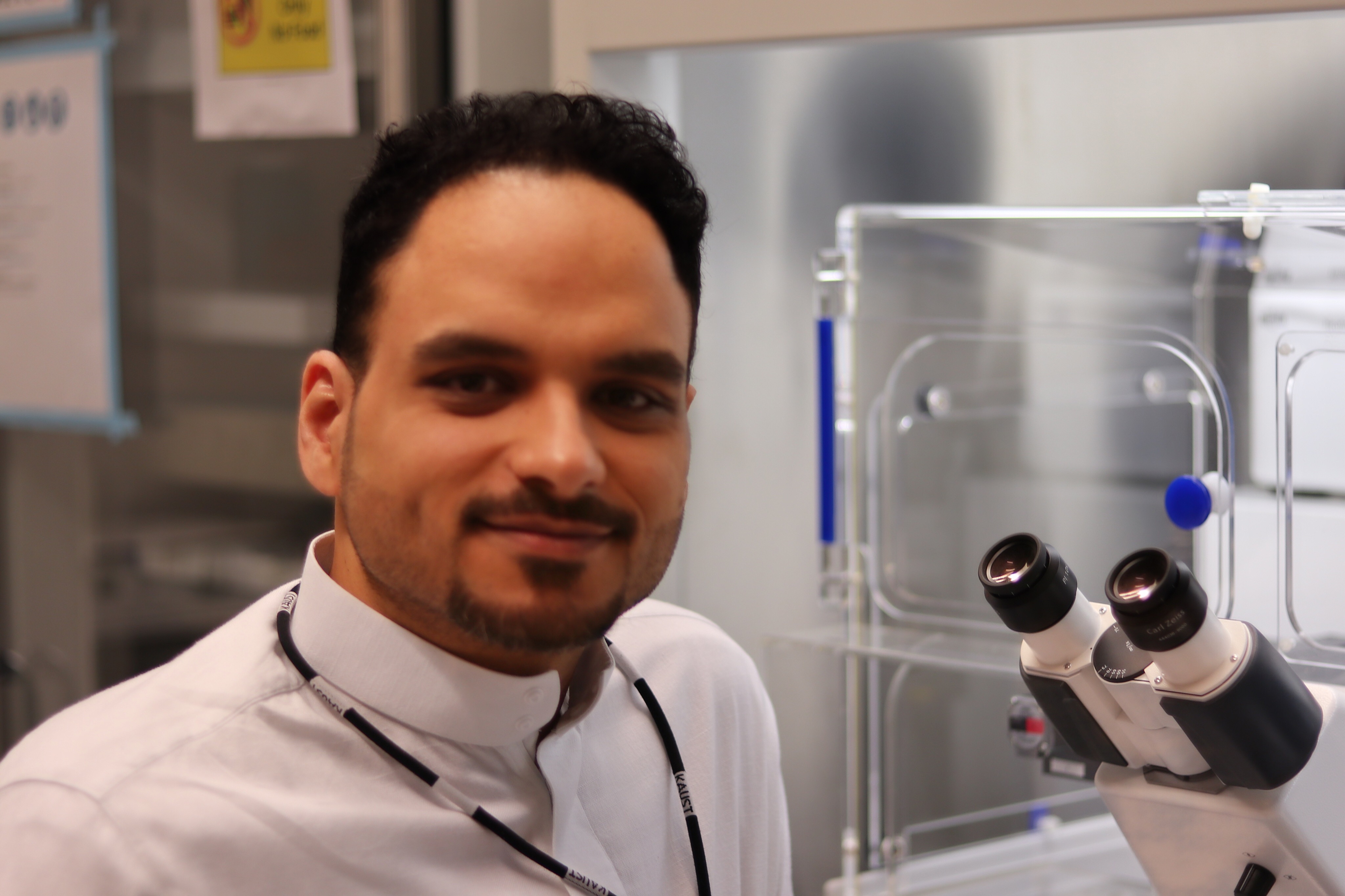

KAUST Ph.D. student Sharif Hala is pictured in the University's labs. Image courtesy of Sharif Hala.
- Taibah University Madinah with Dr. Naif Al Montashiri
- The Saudi Ministry of Health, The General Directorate of Health Affairs Makkah Region Makkah with Dr. Asim Khogeer
- King Fahad Medical Research Center, King Abdulaziz University Jeddah with Dr. Anwar Hashem
- Madinah hospitals - General Hospital, Ohud Hospital, King Fahad Hospital and Maternity and Children Hospital with Dr. Fadwa AlOfi
- Makkah hospitals - Al-Noor Hospital, East Arafat Hospital and Public Health Office with Dr. Khaled AlQithami and Dr. Nashwa Khotani
- Jeddah hospitals - King Abdullah Medical Complex and National Guard Health Affairs with Dr. Afrah AlSomali
"As the coronavirus outbreak [gained momentum], I started up approaching everyone who could help in this matter to get the proper ethical approvals and the paperwork done," said Hala. "As a pathogen genomics lab, it's our duty to be involved hands-on [in tackling] this epidemic."
This means arranging for the transportation of samples from hospitals and quarantine hotels; safely transporting them to KAUST; proceeding to do the viral RNA extraction using optimized kits from the nasopharyngeal swabs and oropharyngeal swab samples; analyzing and quantifying the extracted RNA; identifying if the samples are COVID-19-positive or not; and then proceeding to the protocol of sequencing.
"These samples mean a lot for us because it means we can also help the hospitals in giving them additional feedback on positive and negative samples," Hala explained. "We can also provide a lot of good feedback to the treating doctors, and we can do the transmission studies. I think this will not only allow KAUST to help Saudi Arabia, but it will also be important for the whole world."
Over 600 samples have so far been collected from patients, and the plan is to collect more for a holistic analysis of the COVID-19 disease in Saudi Arabia.
From hospitals to KAUST laboratories
After the typical two days of full-time work needed to generate libraries to prepare for sequencing, the final step is proceeding to sequencing technology machines available to the team in the KAUST Bioscience Core Lab.
"I'm so grateful that the Bioscience Core Lab has opened their doors to do just this part of the work," Pain said. "Without their help, this would not be possible."
In normal times, the MoH would take care of transporting the collected samples, but in these extraordinary times of social distancing and curfews, things are more complicated. Following strict WHO guidelines, Hala developed a very efficient and elaborate process to transport the samples to KAUST. Once he obtained the ethical approvals to collect patient samples from the hospitals, Hala coordinated with medical consultants at the various facilities to make everything possible.
"I give them [full recognition]," he said. "They are the real soldiers who go in and collect the samples from the patients. Myself, I don't really walk into the patient's room; I just make sure that [the sample is] collected and stored okay."
Storing, transporting and ultimately delivering the samples is no small feat. The WHO requires samples to be transported in three covers. The sample is deposited in a tube, which is then placed in a biohazard bag. The bag is then placed inside a box, which is finally secured into a bigger box.
"The big box, which is filled with ice, then gets put inside a bigger biohazard bag that is taped with duct tape to my truck at the back. Basically it's like four enclosures," Hala said.
Due to health and safety reasons, Hala is unable to access the campus and then drive another one to two hours towards his home in Jeddah. On a subsequent collection of 120 samples in Madinah after a spike, he was collecting samples until 4:30 a.m. and ended up meeting Pain at KAUST at 8:30 a.m.
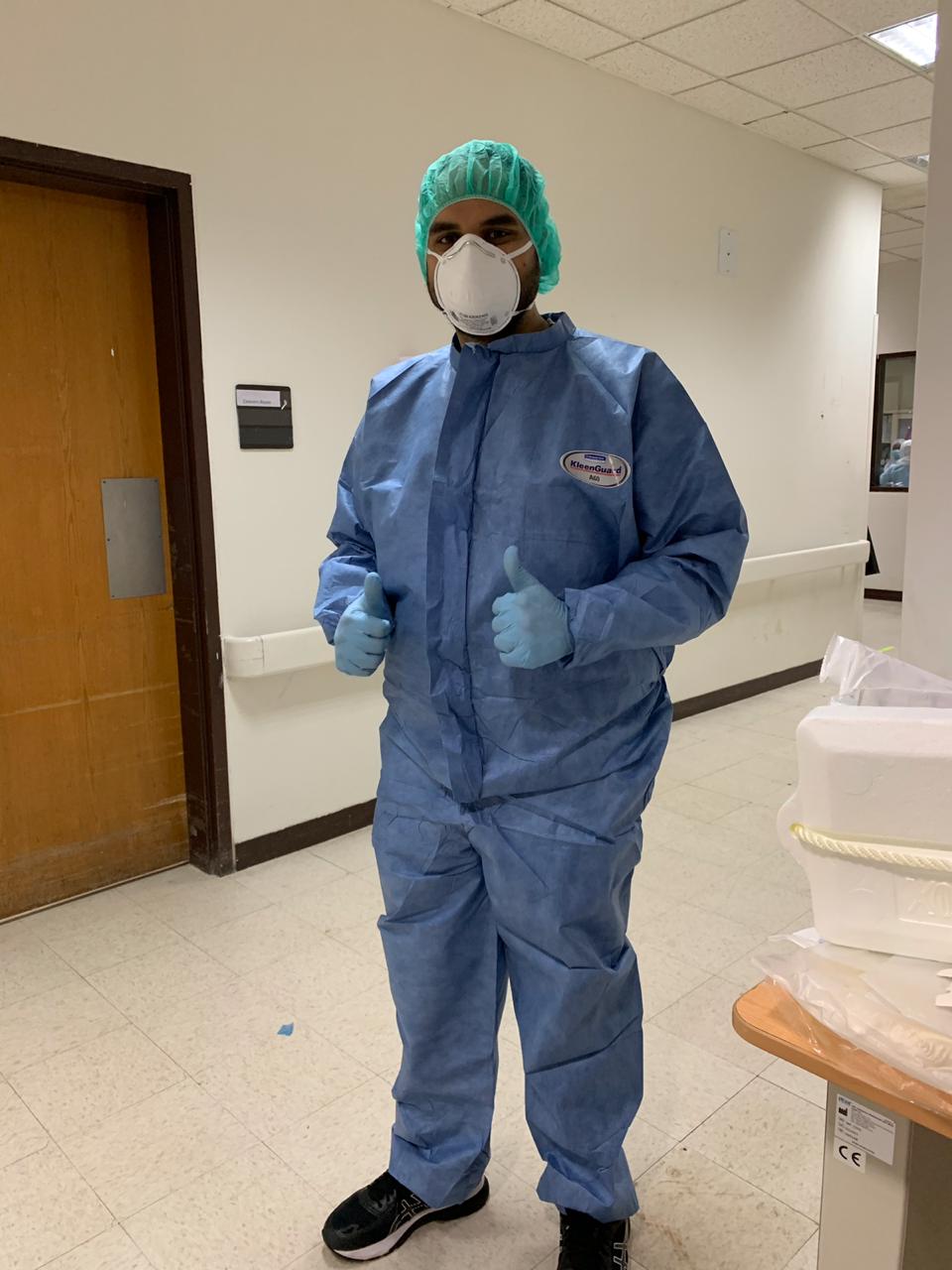

KAUST Ph.D. student Sharif Hala is pictured in one of Saudi Arabia's many Western region medical facilities that he visited to collect samples for the research efforts surrounding COVID-19. Image courtesy of Sharif Hala.
Identifying genetic signatures and cataloging virus strains
One of the critical aspects global researchers are focusing on is understanding how the virus mutates as it gets transmitted from one person to another. By obtaining samples collected at different time points and from a variety of patients, it's possible to identify mutational signatures-or a genetic bar code-that could potentially help to identify the transmission root of a particular strain.
"What we're [now] systematically looking at is the genetic signatures or mutations in these virus strains. [We're] identifying the genomes-basically revealing the genome sequences and then studying the mutations that they carry," Pain said.
It's also essential to be able to have access to a scientific/medical metadata repository associated with samples being sequenced. However, the first step, which the KAUST PGL is currently doing, is getting to know the genetic makeup of the viral strains that the team is sequencing. Eventually they'll have the capacity to compare them with global epidemiological, medical and clinical data sets.
KAUST Professor Arnab Pain works in a University lab with his colleagues. Image courtesy of Arnab Pain.
A catch-up game
Scientists only got to know the SARS-CoV-2 genome less than a month ago. There are now over 8,700 sequences available, so it's possible to utilize bioinformatics power to understand the novel genome and its modifications. Learning how and where it's mutating, which genes are changing and which mutations have a more significant impact on the biology are of top importance.
"We like to understand how do these COVID cases-the viruses that are circulating in the population in Saudi Arabia-are linked to each other," Pain explained. "Also, more importantly, how are they linked to the rest of the world, because most of the initial cases in Saudi Arabia [were] imported by international travelers."
"If you systematically sequence all these genomes and identify the genetic code of each of the strains and variation in that code over time, then you can systematically track what kind of mutations they carry in the Saudi strains," he continued. "Then it's possible to compare them with other samples from the Middle East and the rest of the world."
Defining the barcode and tracking the strains in an epidemiological setting allows scientists to understand how the viruses are moving in the population and how they're changing. It also enables them to get a sense as to whether they're becoming more transmissible or more virulent or less so.
"We're doing a catch-up game at the moment as scientists," Pain said. "[Finding answers to more questions] will be possible when we have an even greater number of genomes available, and we are just doing our bit to achieve that goal."






