A group of researchers have demonstrated that, from seven methods commonly used to test for viruses in untreated wastewater, an adsorption-extraction technique can most efficiently detect SARS-CoV-2. This gives us another tool to detect the presence and spread of the COVID-19 pandemic.
Tracking the spread of the COVID-19 pandemic is currently conducted by testing nasal swabs or saliva samples. Tools and techniques to track the spread of the pandemic by other means would be very beneficial; wastewater monitoring is a method that would allow us to monitor the spread of the pandemic at a much larger scale. This is not a new technique, and has been used for detecting non-enveloped viruses, but a conventional method for enveloped viruses such as SARS-CoV-2 had not been developed.
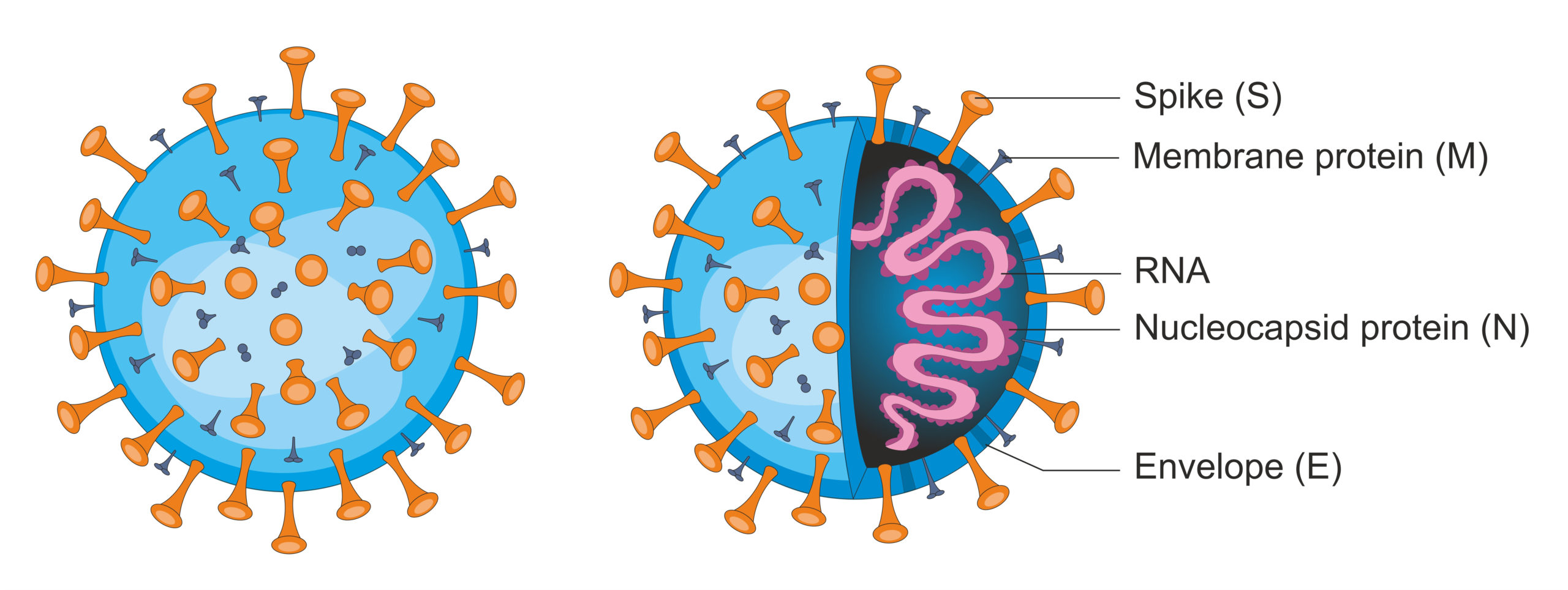
SARS-CoV-2. All viruses have genetic material (DNA or RNA) surrounded by a protein coat. Some viruses - the enveloped viruses - also possess an outer coat of lipids. (Soleil Nordic/Shutterstock)
In the current work, co-authored by Assistant Professor Masaaki Kitajima from the Water Quality Control Engineering Laboratory at Hokkaido University, scientists report a fast, economical method to concentrate coronavirus in untreated wastewater. Murine hepatitis virus (MHV), a type of enveloped virus, is closely related to SARS-CoV-2 but does not affect humans, and is thus safe to use for testing the feasibility of the method. The study was published in Science of the Total Environment.
The scientists obtained MHV from mice feces and introduced it into samples of untreated wastewater collected from Brisbane, Australia. They attempted to recover and concentrate the MHV from these samples by seven different methods which are commonly used to test for non-enveloped viruses. The amount of recovered MHV was determined by a method called reverse transcription-quantitative PCR, where the RNA of the virus extracted, converted to DNA, the DNA is repeatedly duplicated, and the increase in amount of DNA is measured throughout the process.
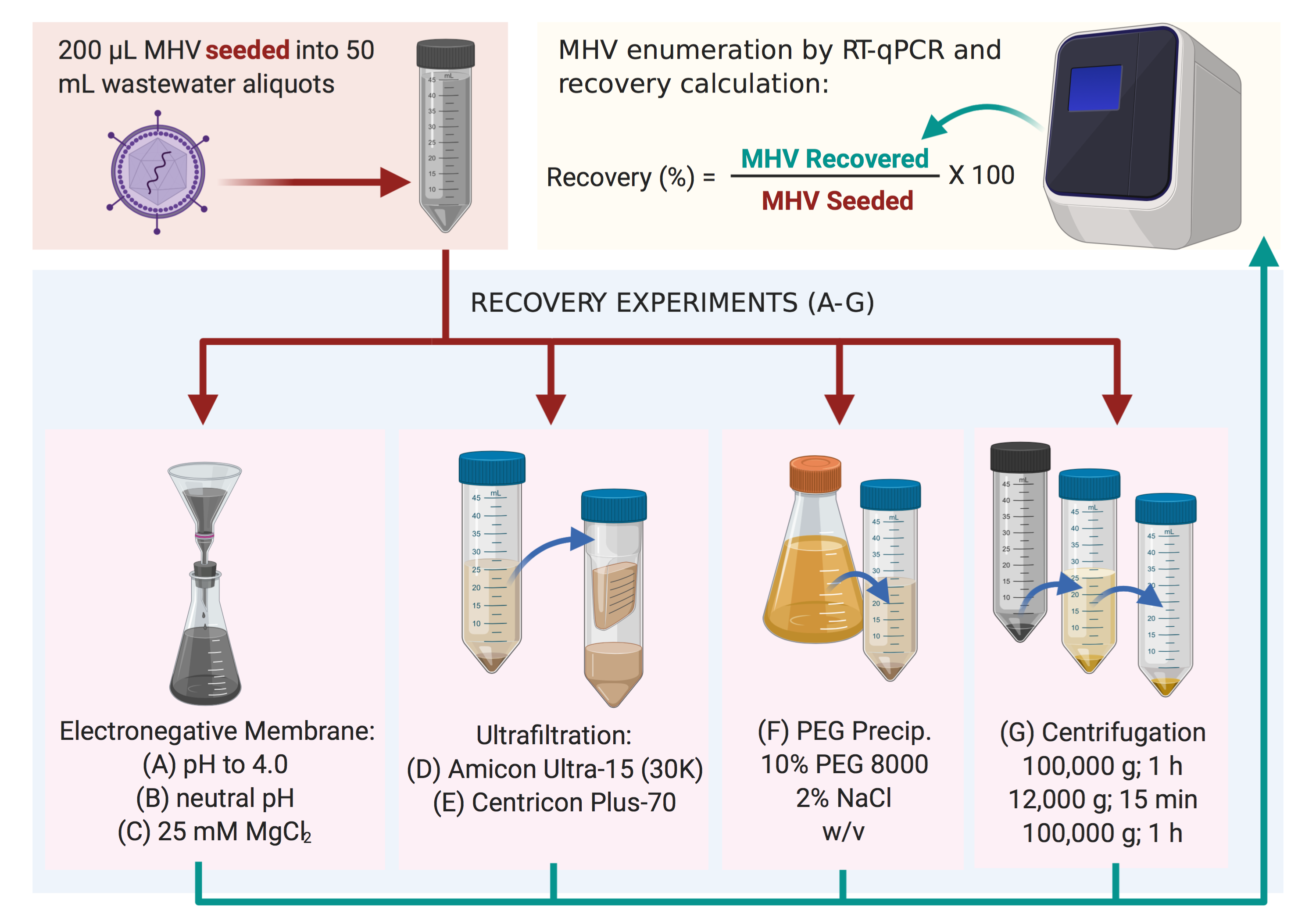
Methods used to recover MHV in this study. The most successful was method (C), followed by method (B) (Warish Ahmed et al., Science of The Total Environment, June 5, 2020)
The recovery was highest in the method that involved treating the sample with magnesium chloride and then filtering out the virus on a negatively-charged membrane; the second highest recovery was by a similar method without magnesium chloride. The advantages of these methods include an initial processing time of under 1 hour and the need only for cheap, widely available equipment and reagents. There are also drawbacks, such as the clogging of the filters that may increase processing time. However, to date, the need for reverse transcription-qPCR for the detection of the virus is unavoidable.
The next step would be to test this method in samples collected from areas where the pandemic is prevalent. There are two objectives: one is to show that the technique can be used for SARS-CoV-2, and the other is to show that the test can be used on samples from outside the lab.
"I hope this research contributes to the establishment of a standard protocol for the detection of SARS-CoV-2 in wastewater," says Assistant Professor Kitajima, "and this, in turn, accelerates investigations to enhance our understanding of COVID-19 epidemiology through wastewater surveillance." He is currently involved in a number of studies related to applying wastewater-based epidemiology to tracking the spread of the COVID-19 pandemic, and has collaborated with a number of scientists and research groups across the world in this endeavor.

Warish Ahmed, CSIRO Land and Water (left) and Masaaki Kitajima, Hokkaido University (right) (Photos provided by Masaaki Kitajima).
Original article:
Warish Ahmed et al., Comparison of Virus Concentration Methods for the RT-qPCR-Based Recovery of Murine Hepatitis Virus, a Surrogate for SARS-CoV-2 from Untreated Wastewater. Science of The Total Environment. June 5, 2020.
DOI: 10.1016/j.scitotenv.2020.139960.
Funding:
This study was funded by the CSIRO Land and Water, Australia; a US National Science Foundation grant (OCE-1566562); and, the Japan Science and Technology Agency (JST) Mirai Program (JPMJMI18DB).






