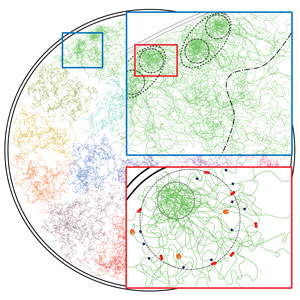
 Figure 1: A schematic depiction of the architecture of chromosomes found by a technique developed by RIKEN researchers that analyzes data derived with high-throughput chromosome conformation capture (Hi-C) experiments. New structures known as enclaves are indicated by the broken black lines in the two insets.
Figure 1: A schematic depiction of the architecture of chromosomes found by a technique developed by RIKEN researchers that analyzes data derived with high-throughput chromosome conformation capture (Hi-C) experiments. New structures known as enclaves are indicated by the broken black lines in the two insets.
Reproduced from Ref. 1 and licensed under CC BY 4.0 © 2020 V. Kumar et al.
A technique developed by three RIKEN researchers can identify interaction patterns within human chromosomes that conventional methods miss1. It will help produce new maps of our chromosomes and uncover the complex interactions in them.
The cells in our bodies vary greatly in terms of shape and functions despite containing the same genetic information. One cause of this variety is the different ways genetic material is packaged in the nuclei of cells, which gives rise to the expression of different genes.
A powerful technique for analyzing the interactions between different regions of a chromosome is high-throughput chromosome conformation capture (Hi-C). The application of different clustering methods to Hi-C data has led to the identification of two levels of chromosomal structure: compartments on a large scale and topologically associating domains (TADs) on a much smaller scale.
"These two types of structures exist within the same chromosome and yet are detected using very different clustering techniques," says Vipin Kumar, who was at the RIKEN Center for Biosystems Dynamics Research at the time of the study. "That was the motivation behind our study-to detect these different scales of organization with the same technique."
Now, Kumar, Simon Leclerc and Yuichi Taniguchi have developed a more general version of the Hi-C data clustering technique, which they call BHi-Cect. It employs an iterative top-down approach. "Starting with the whole chromosome, BHi-Cect progressively separates it into its most likely clusters, beginning with the biggest ones and then looking for smaller ones," explains Kumar.
 Vipin Kumar (second row, third from right), Simon Leclerc (second row, second from right) and Yuichi Taniguchi (second row, middle) with their group. They have developed BHi-Cect, a top-down algorithm for finding clusters in chromosomes, and used it to identify structures called enclaves.
Vipin Kumar (second row, third from right), Simon Leclerc (second row, second from right) and Yuichi Taniguchi (second row, middle) with their group. They have developed BHi-Cect, a top-down algorithm for finding clusters in chromosomes, and used it to identify structures called enclaves.
© 2020 RIKEN Center for Biosystems Dynamics Research
"Unlike previous methods, BHi-Cect does not assume specific criteria to identify different structural features," says Taniguchi. "BHi-Cect allows us to identify many global and local structural features, including compartments, TADs and looping structures, and to associate them with a hierarchical tree-it's a completely different approach."
Kumar, who is now at the University of Oslo, agrees: "We don't need to choose between looking for local or global structures; this technique gives you everything at once-that's the big difference."
Using BHi-Cect, the trio identified structures that they termed enclaves (Fig. 1). These include both compartments and TADs, but also other interactions that don't fall into either category.
"We applied a very general definition: there is more interaction within an enclave than with the rest of the chromosome," explains Kumar. "Consequently, our technique was able to detect much more-subtle interaction patterns in Hi-C data. I think this reflects the fact that chromosome structures are much more complex than previously thought."






