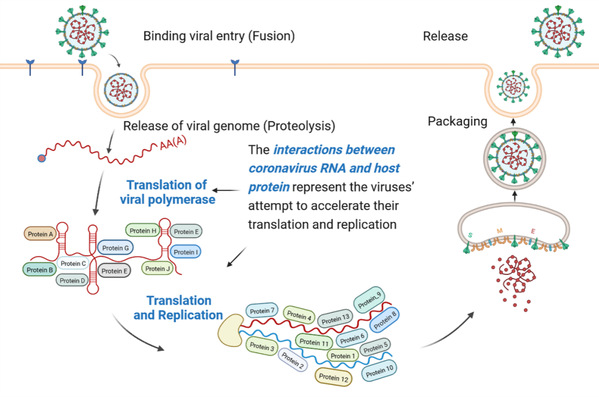
Coronavirus has brought about three massive outbreaks in the past two decades. Each step of its life cycle invariably depends on the interactions among virus and host molecules. The interaction between virus RNA and host protein (IVRHP) is unique compared to other virus-host molecular interactions, and has emerged to be a very hot topic in recent studies (Cell. 184: 2394-2411.e16, 2021; Cell Res. 32: 9-23, 2022). These studies provide essential information for a deeper understanding of IVRHP, which represents not only an attempt by viruses to promote their translation/replication, but also the host's endeavor to combat viral pathogenicity. In other words, there is an urgent need to have a panorama of coronavirus RNA-Host protein interactions, which will then aid in the discovery of new antiviral therapies.
On October 06, 2022, Prof. ZHU Feng from College of Pharmaceutical Sciences in Zhejiang University, Prof. HAN Lianyi from College of Life Sciences in Fudan University and Prof. Lin TAO from College of Pharmaceutical Sciences in Hangzhou Normal University published an article entitled "CovInter: Interaction Data between Coronavirus RNAs and Host Proteins" in the journal of Nucleic Acids Research. Just as Prof. ZHU says, "Given the devastating and persistent threat of coronaviruses, CovInter is highly expected to fill the information gap in the whole process of the 'molecular arms race' between coronaviruses and their hosts, which will therefore facilitate the identification of new therapeutic targets for drug discovery/repurposing."

Prof. ZHU Feng (the seventh from the right on the second line) and his team.
In this study, Prof. ZHU's research team focused on constructing the panorama of coronavirus RNA-host protein interactions, and a knowledge base named CovInter was developed, for the first time, to provide the interaction data between virus RNA and host protein. This knowledge base comprehensively described the experimentally verified function of thousands of host proteins in viral infection and their ability to sneak past the immunity. It also quantitatively demonstrated the differential expression patterns and phosphorylation sites (before and after infection) of these key proteins. Furthermore, to achieve an in-depth insight into the host response pathway activated by host proteins during infection, hundreds of infection-associated pathways were also systematically identified.
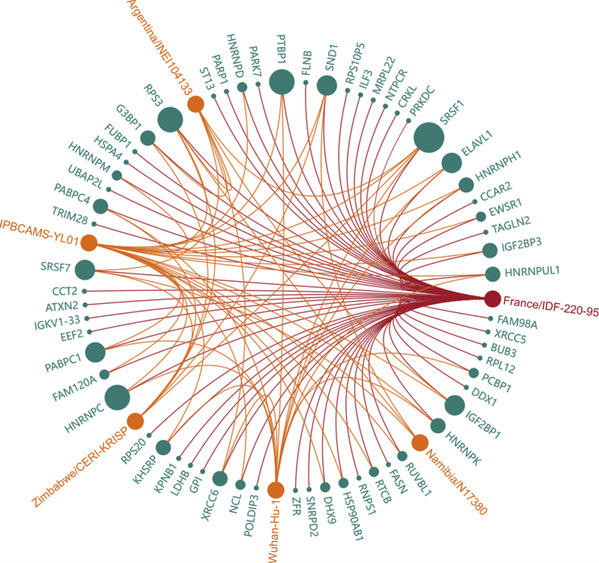
Moreover, due to the rapid variation in coronavirus RNA, significant gain/loss of interaction (especially IVRHPs) has been frequently reported, which is highly expected to lead to substantial changes in the rate of both virus transmission and case fatality.Thus, it is key to have an in-depth understanding of the level of the conservation among the IVRHPs of various virus variants/strains.Therefore, in this study, Prof. ZHU's team systematically mapped the IVRHP interaction network for each virus RNA and then demonstrated their level of conservation among virus strains (as shown in the figure below).The red circle represents a particular RNA, green circles indicates different host proteins, orange circles denotes other RNAs (from other strains of SARS-CoV-2) which interacted with the same host protein as that of the studied RNA. The diameter of green circle indicated the number of virus' RNAs that could interact with the host protein. The larger the diameter of a protein is, the more virus RNAs this protein interacts with. The profile reflects the molecule's role in the life cycle of the coronavirus. All in all, this study identified that the conserved interactions between coronavirus RNA and host proteins were highly variable.
Source: College of Pharmaceutical Sciences, Zhejiang University






