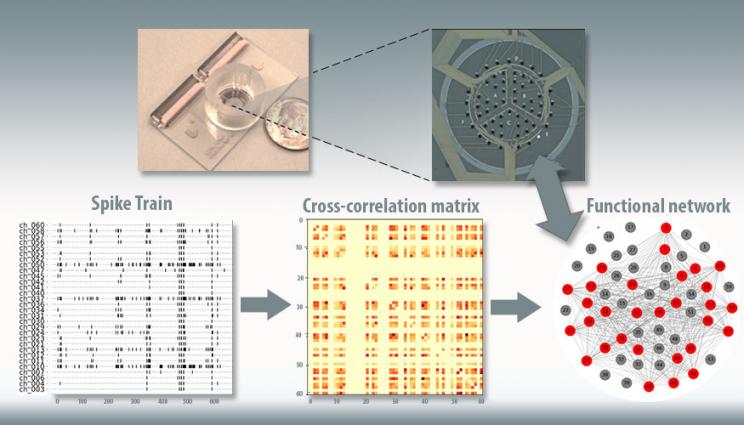
This figure depicts Lawrence Livermore National Laboratory's "brain-on-a-chip" device (top). Electrical recordings from the chip (lower left), taken from seeded neurons, are used to model correlations between electrodes (center), and these correlations are used to create a model of the network structure in the chip (lower right).
For the past several years, Lawrence Livermore National Laboratory (LLNL) scientists and engineers have made significant progress in development of a three-dimensional "brain-on-a-chip" device capable of recording neural activity of human brain cell cultures grown outside the body.
Now, LLNL researchers have a way to computationally model the activity and structures of neuronal communities as they grow and mature on the device over time, a development that could aid scientists in finding countermeasures to toxins or disorders affecting the brain, such as epilepsy or traumatic brain injury.
As reported recently in the journal PLOS Computational Biology, an LLNL team has developed a statistical model for analyzing the structures of neuronal networks that form among brain cells seeded on in vitro brain-on-a-chip devices. While other groups have modeled basic statistics from snapshots of neural activity, LLNL's approach is unique in that it can model the temporal dynamics of neuronal cultures - the evolution of those neural network changes over time. With it, researchers can learn about neural community structure, how the community evolves and how the structures vary across experimental conditions. Although this current work was developed for 2D brain-on-a-chip data, the process can be readily adapted to LLNL's 3D brain-on-a-chip.
"We have the hardware but there's still a gap," said lead author Jose Cadena. "To really make use of this device, we need statistical and computational modeling tools. Here we present a method to analyze the data that we collect from the brain-on-a-chip. The significance of this model is that it helps us bridge the gap. Once we have the device, we need the tools to make sense out of the data we get from it."
Using thin-film multi-electrode arrays (MEAs) engineered into the brain-on-a-chip device, researchers have successfully captured and collected the electrical signals produced by neuronal networks as they communicate. With this data as teaching tools, the team combined stochastic block models that are standard in graph theory with a probabilistic model called Gaussian process that includes a machine learning component, to create a temporal stochastic block model (T-SBM).
The model was applied to three datasets; culture complexity, extracellular matrix (ECM) - the protein coating the cells are grown on - and neurons from different brain regions. In the first experiment, researchers looked at data on cultures containing only neuronal cells versus cultures that had neurons mixed with other types of brain cells, closer to what one would find in a human brain. Researchers found what they would expect, that in more complex cultures that contained other cell types, the networks that develop are more complex and communities get more intricate over time. For the second study with ECM, the model analyzed neurons grown in three different kinds of tissue-like proteins, finding that the coating in which these neurons are grown on the device has little effect on the growth of neural cultures. The datasets for the first two studies were generated through brain-on-a-chip experiments performed at LLNL and led by LLNL researchers Doris Lam and Heather Enright.
"We knew from our experiments that numerous neuronal networks have been formed, but now with this statistical model we can identify, distinguish and visualize each network on the brain-on-a-chip device and monitor how these networks change across experimental conditions," Lam said.
In the last study, researchers observed differences in the networks in cortical and hippocampal cultures, showing a much higher level of synchronized neural activity in hippocampal cultures. Taken together, researchers said the results show that the temporal model is capable of accurately capturing the growth and differences in network structure over time and that cells are able to grow networks on a chip-based device as described in neuroscience literature.
"These experiments show we can represent what we know happens in the human brain on a smaller scale," Cadena said. "It's both a validation of the brain-on-a-chip and of the computational tools to analyze the data we collect from these devices. The technology is still brand new, there aren't many of these devices; having these computational tools to be able to extract knowledge is important moving forward."
The ability to model changes in neural connections over time and establish baseline normal neural activity could help researchers use the brain-on-a-chip device to study the effects of interventions such as pharmaceutical drugs for conditions that cause changes in network structures to the brain, such as exposure to toxins, disorders such as epilepsy or brain injuries. Researchers could develop a healthy brain on a chip, induce an epileptic attack or introduce the toxin and then model the effect of the intervention to revert to the baseline state.
"It's essential to have this kind of computational model. As we begin to generate large amounts of human-relevant data, we ultimately want to use that data to inform a predictive model. This allows us to have a firm understanding of the fundamental states of the neuronal networks and how they're perturbed by physical, chemical or biological insults," said principal investigator Nick Fischer. "There's only so much data we can collect on a brain-on-a-chip device, and so to truly achieve human relevance, we'll need to bridge that gap using computational models. This is a stepping-stone in developing these sorts of models, both to understand the data that we're generating from these complex brain-on-a-chip systems as well as working toward this kind of predictive nature."
The work was funded by the Laboratory Directed Research and Development (LDRD) program and was one of the final steps of a Lab Strategic initiative (SI) to develop and evaluate neuronal networks on chip-based devices. As part of this project, the team also optimized the biological and engineering parameters for 3D neuronal cultures to better understand how architecture, cellular complexity and 3D scaffolding can be tuned to model disease states with higher fidelity than currently possible.
With a validated device in place, the Lab team is pursuing funding from external sponsors to use the 3D brain-on-a-chip to screen therapeutic compounds and to develop human-relevant models of neuronal cultures for diseases and disorders such as traumatic brain injury, in an effort to find ways of re-establishing normal brain function in TBI patients.
"All of the work we've done under this SI underscores the Lab's commitment and strategic investment into developing these organ-on-a-chip platforms," Fischer said. "We're coming to a place where we understand how to properly design and implement these platforms, especially the brain-on-a-chip, so we can apply them to answer questions that are relevant to national security as well as to human health.
"It's a long road to develop these really complex systems and to tailor them for the specific applications of interest to the Lab and the broader research community," he continued. "This isn't something that could come out of a single group: it really requires the kind of multidisciplinary team that you find at a national lab that helps bring something like this to fulfillment."
Co-authors on the paper included research engineer and deputy director for the Lab's Center for Bioengineering Elizabeth Wheeler and former Lab computational engineer Ana Paula Sales.






