The genomes of phages - viruses that infect bacteria - are largely composed of "dark matter": genes that encode proteins whose functions remain unknown. Less than four years ago, a team led by Prof. Rotem Sorek at the Weizmann Institute of Science identified a new type of protein within this viral dark matter and dubbed it a "sponge." Viral sponge proteins are porous and specialize in trapping molecules within deep pockets - much like a sponge that absorbs water. For phages, however, this sponge serves as a weapon: It traps communication molecules that are essential to bacterial immune systems, allowing the phage to take control of the bacterium and multiply inside it unhindered.

Until recently, very few sponge proteins had been found. Their genetic sequences differ greatly from one another, making them difficult to detect. Now, using an innovative research approach that combines artificial intelligence with experimental biology, researchers in Sorek's lab have uncovered new families of sponge proteins that disrupt immune communication in bacteria. The findings, published in Science , reveal how viruses silence the immune system's alarm signals, and shed light on the importance of communication disruption in the billion-year-long war between viruses and bacteria.
In the new study, the researchers examined the structures of sponge proteins identified so far and noticed a recurring architectural pattern that could be used to discover new proteins of this type. "They are all small, composed of several identical subunits and contain deep pockets," explains Sorek. "These pockets carry a positive electrical charge, allowing them to absorb immune alarm molecules, which are typically negatively charged."
""A student's keen observation helped us identify a new family of sponge proteins - a reminder that human insight still matters in the age of AI"
Insights like these used to have limited practical value, but the AI revolution has changed that. "We realized that with advanced AI tools such as Google's AlphaFold, we could scan an enormous number of proteins and search for those with positively charged pockets capable of trapping immune molecules," says Dr. Nitzan Tal, who led the new study in Sorek's lab. "This allowed us to reveal new functions of phage proteins based solely on their structure."
The scientists scanned a database of 32 million genes encoding phage proteins, from 2 million phage genomes, and used AlphaFold to predict their three-dimensional structures. "We found more than 120 candidates whose structures matched our criteria, and moved on to experimental testing," says Tal.
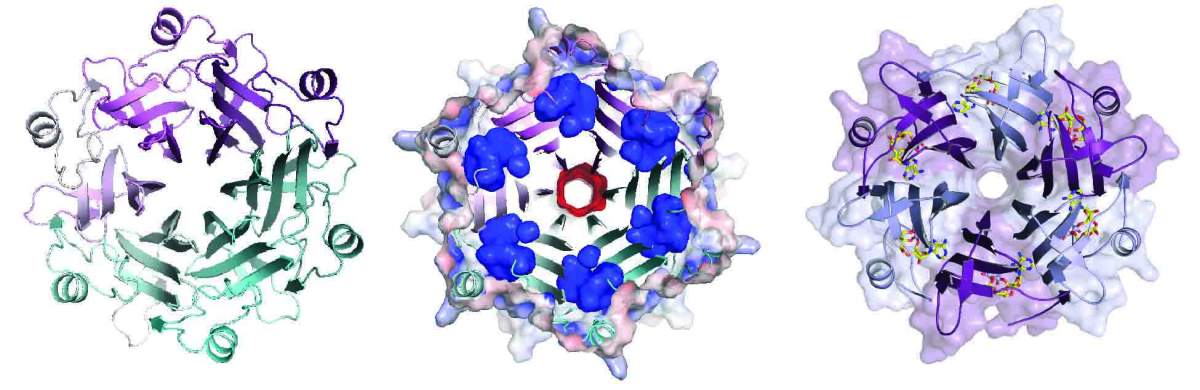
The researchers then tested the effectiveness of each candidate against five bacterial immune systems, using a new method developed by research student Jeremy Garb in Sorek's lab. The approach enabled the team to perform all the tests simultaneously rather than conducting hundreds of separate experiments. These experiments revealed a new family of sponge proteins that the researchers named Lockin. The AI model predicted that these proteins should consist of six identical subunits arranged in a circular structure resembling flower petals. In collaboration with Prof. Philip J. Kranzusch's team at the Dana-Farber Cancer Institute in Boston, the researchers determined the structure of one family member using X-ray crystallography, confirming the prediction and deciphering exactly how the immune alarm molecule is captured.
"The huge database of viral proteins we analyzed was mostly obtained from sequencing environmental DNA samples that include a large mixture of phages," says Sorek. "This allowed us to discover the Lockin proteins, which appeared in hundreds of phages that have never been isolated in the lab."
Along with AI-based predictions, the researchers used additional innovative strategies. "Romi Hadary, another research student in my lab, noticed that genes that encoded known sponge proteins tend to be fused together in phage genomes," explains Sorek. "This insight allowed us to identify an additional family of sponge proteins, called Sequestin, based on the fact that their genes are fused to those of known sponges. It goes to show that, even in the age of artificial intelligence, there is still great value in the keen observations of human scientists."
Yet another protein family discovered in the study, called Acb5, initially puzzled the researchers. "These proteins were very similar to sponge proteins, but we discovered that they not only trap alarm molecules - they also cut them," says Tal. "This was surprising because they didn't have the structural features typical of molecular cutting tools. This discovery shows how systematic structural scanning can overturn previous scientific assumptions."
The protein families identified in this study appear in the genomes of thousands of different phages in nature. The researchers also found that a single phage can carry a broad arsenal of sponges and enzymes that neutralize immune alarm molecules. Together, these findings show that proteins disrupting immune communication give phages a significant advantage in their arms race with bacteria.
"It's not yet known whether viruses that infect plants, animals and humans also use sponge proteins, but the computational and experimental approach we developed makes it possible to test this," adds Sorek. "If they do, sponge proteins could become targets for the development of antiviral therapies in the future. Our discovery method doesn't require prior knowledge of protein function, and it doesn't rely on spotting similarities in genetic sequences or on growing viruses in the lab. It is therefore a powerful tool for uncovering additional immune-related proteins that share structural patterns."
Science Numbers
The function of approximately 70 percent of phage genes is still unknown.
Also participating in the study were: Dr. Ilya Osterman, Dr. Gil Amitai, Erez Yirmiya, Dr. Nathalie Béchon, Dr. Dina Hochhauser and Barak Madhala from Weizmann's Molecular Genetics Department; Renee B. Chang and Miguel López Rivera from the Dana-Farber Cancer Institute, Boston, MA; Roy Jacobson from Weizmann's Plant and Environmental Sciences Department; Dr. Moshe Goldsmith from Weizmann's Biomolecular Sciences Department; and Dr. Tanita Wein from Weizmann's Systems Immunology Department.
Prof. Rotem Sorek's research is supported by Magnus Konow in honor of his mother Olga Konow Rappaport.






