Evolution can promote novelty by keeping gene expression in check
Changes in the genes that control development can potentially make large contributions to evolution by generating new morphologies in plants and animals. However, because developmental genes frequently influence many different processes, changes to their expression carry a risk of "collateral damage". Scientists at the Max Planck Institute for Plant Breeding Research in Cologne, and collaborators, have now shown how gene self-repression can reduce the potential side effects of novel gene expression so that new forms can evolve. This self-regulation occurs via a distinctive molecular mechanism employing small regions of genomic DNA called low-affinity transcription factor binding sites.
Suppose a bird develops a modified wing shape, which makes flying easier and could be beneficial to its survival. If this gene change also altered the bird's color, making it less attractive to mates, then the advantageous wing-shape modification would be unlikely to persist. So, how then does nature balance the potential for novelty, with the risk of side effects that may prevent novelty from arising? Using the evolution of leaf shape as an example, an international team led by Director Miltos Tsiantis has provided fresh insight into this question.
This new study was done in the hairy bittercress, a small weed that the Tsiantis group has developed into a model system for understanding evolution of plant form. It builds on previous work from the group in which a gene called RCO was found to have driven leaf shape diversification in mustard plants by acquiring a novel expression pattern.
RCO encodes a transcription factor, a type of protein that can turn other genes on or off, and RCO's new expression pattern resulted in the emergence of the more complex leaf shapes found in bittercress. The researchers have now shown that this change in gene expression was accompanied by RCO acquiring the ability to repress its own activity. Mike Levine, Director of the Lewis-Sigler Institute for Integrative Genomics at Princeton University who was not involved in the study, finds this particular insight "very compelling". As the self-repression of RCO "limits the scope of its activity", Levine explains, it "thereby blocks potentially deleterious influences on cell development and function".
Stimulating cytokinin
As a next step, the scientists identified the genes targeted by RCO, and found that many of them are responsible for coordinating local levels of cytokinin - a widely acting plant hormone known to affect cell growth. Importantly, when the self-regulation of RCO is modified, RCO stimulates cytokinin excessively and leaf shape is altered in ways that can negatively affect plant fitness. This finding confirms the idea that self-repression of RCO could be essential for the persistence of RCO-induced novel leaf morphologies.
What's particularly interesting is that this self-repression of RCO occurs in a very distinctive way. The scientists discovered that it is based on many weak interactions between the RCO protein and RCO regulatory DNA at low-affinity binding sites. "This finding is exciting", explains Tsiantis, "because low-affinity binding sites can evolve relatively quickly, thus offering an easy way for evolution to keep changes in gene expression in check, by lowering a regulator's expression".
Soft repression
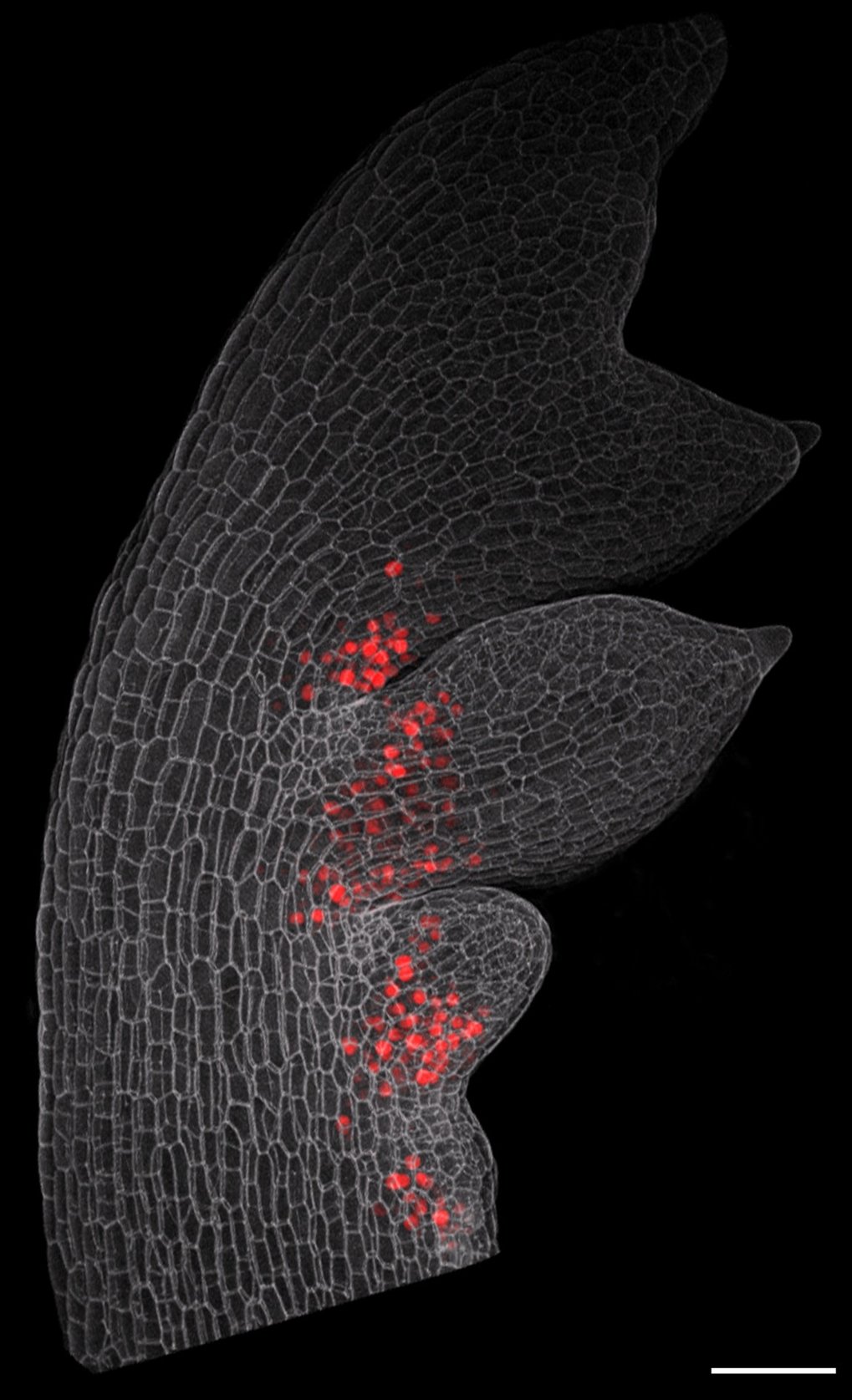
Indeed, this latest work from Tsiantis's team directly demonstrates that low-affinity transcription factor binding sites can play a major role in the generation of morphological novelty. By providing a tool to "softly" repress RCO expression, these sites dampen the effects of RCO expression changes and allow cytokinin levels to be fine-tuned. This in turn promotes the appearance of more complex leaf shapes, e.g., by precisely regulating the outgrowth of lobes or leaflets along the margins of developing leaves.
These results will stimulate further efforts to understand the influence of low-affinity transcription factor binding sites on development, diversity and disease. For example, there is increasing awareness that changes in the regulation of developmental genes are a major contributor to human disease, and that other regulatory changes can reduce disease severity or protect individuals who carry disease variants. While the specific DNA sequences underlying these effects are often unknown, this latest work highlights low-affinity transcription factor binding sites as excellent candidate regions for identifying causal sequences of disease susceptibility, and for understanding variation in trait diversity more broadly in complex eukaryotes.






