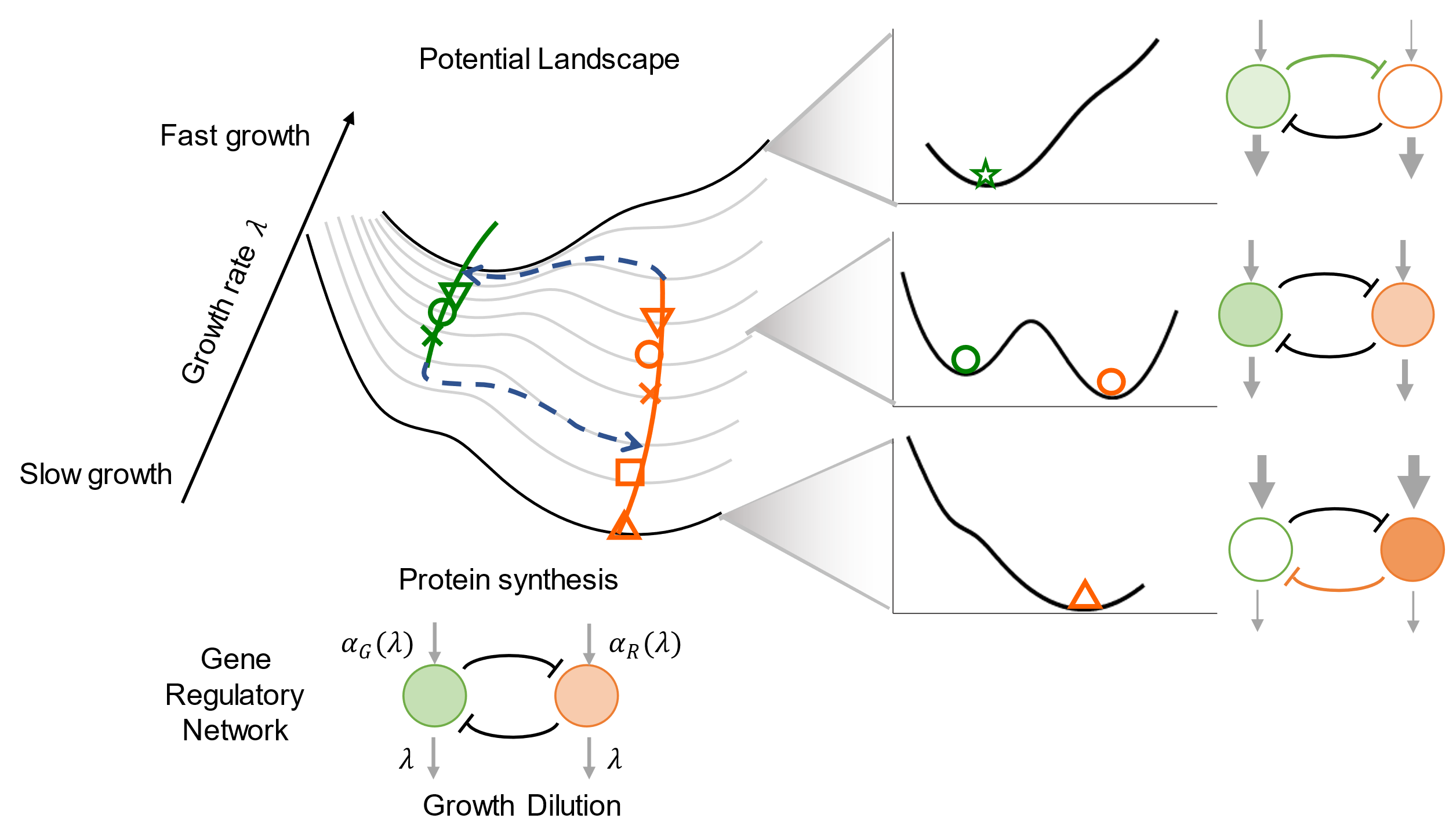
Genes and the regulation relationships among them create complex networks that determine cell differentiation trajectories. However, we still cannot understand and predict the cell-fate-decision process using network topology in a bottom-up manner.
Recently, a research group led by Prof. FU Xiongfei from the Shenzhen Institute of Advanced Technology (SIAT) of the Chinese Academy of Sciences (CAS) has revealed how the global regulation factor, cellular growth rate, reshapes the cell-fate-decision landscape.
The study was published in Nature Chemical Biology on March 23.
The researchers utilized a classical synthetic genetic circuit, the toggle switch, as a toy model to study the extent to which cellular growth rate affects the phenotypic states of the cell. They found that the gene expression response to growth-rate variations was unbalanced, resulting in a growth-rate-dependent phase transition between the available phenotypic states of the toggle.
They also analyzed the bistability of the toggle under various growth conditions, revealing that the steady states bifurcate at a critical value of the cell's growth rate. They quantified the gene expression capacities at different growth rates and proposed a mathematical model that elucidates the bifurcation arising from the unbalanced relationship between gene expression and growth rates.
"To further verify the model's capabilities, we constructed toggles with tuned intrinsic parameters and successfully predicted their steady-state properties," said Prof. FU, corresponding author of the study.
This global regulation mechanism doesn't rely on specific regulatory factors. It is ubiquitous in any genes in the cell-fate-decision network and can emerge or program a new gene expression pattern with a spatiotemporal order.
"It is hard to disentangle the global factors, such as growth rate, from the cell-fate-decision processes," said Prof. FU, noting that the growth rates can be varied as the results of the fate decision. "It's a chicken and egg decision."
With the aid of quantitative synthetic biology, however, researchers can reconstruct orthogonal networks that insulate host genetic networks. In this way, they can evaluate how growth rate affects gene expression and how they promote the bifurcation of cell phenotypic states.