A multinational research team led by researchers at Institute of Science Tokyo, RIKEN, and the University of Toronto has revealed how a tryptophan-rich allosteric communication network regulates receptor dynamics and activation of the human adenosine A2A receptor (A2AR), a major G protein-coupled receptor (GPCR) drug target. By integrating experimental functional assays and residue-specific NMR with molecular simulations and fast allostery-prediction algorithms based on rigidity theory, the team mapped long-range allosteric communication pathways linking the ligand-binding pocket to the intracellular G protein-coupling machinery and identified a central role for tryptophan residues along these pathways. The study also clarifies the functional role of the receptor's conserved sodium-binding pocket, showing that sodium egress strongly promotes activation-related conformational states, including a precoupled state that likely prepares the receptor for productive G protein interaction. These findings deepen our understanding of GPCR activation and allostery, and may support future development of allosteric GPCR drugs.
Beyond the specific mechanism, this work addresses a major bottleneck for AI in structural biology: recent advances such as AlphaFold have transformed prediction of static protein structures, but AI still cannot reliably predict the dynamics and allosteric communication that determine function, signaling, and drug response. To help close this gap, the researchers developed and applied fast computational methods for probing allosteric and dynamic regulation in protein structures and anchored these predictions with experimental NMR validation. The resulting experimentally validated, computationally generated data on allostery and dynamics-and a scalable approach to extend these datasets across diverse receptors and conditions-provide scarce, high-value training and benchmarking data for next-generation AI models aimed at predicting protein function beyond static structure, accelerating future AI-driven prediction of protein function and the design of selective GPCR therapeutics.
Background
GPCRs are the largest family of membrane receptors in humans and among the most important targets in modern medicine. Although many approved drugs act on GPCRs, designing therapies that are both potent and highly selective remains challenging because GPCR function depends not only on static structures, but also on dynamic conformational changes controlled by long-range allosteric communication within the receptor and between the receptor and its signaling partners. AI structure prediction has advanced rapidly, but predicting dynamics and allostery remains a key challenge for understanding protein function.
The research team focused on the human adenosine A2A receptor (A2AR), a prototypical class A GPCR widely distributed in the nervous system, platelets, immune cells, lungs, heart, and vasculature. A2AR can engage multiple G proteins: beyond its cognate GS signaling, it has also been reported to engage other G proteins, highlighting the broader challenge of understanding G protein selectivity and efficacy in GPCR signaling.
A2AR-targeting drugs have been developed for diverse indications, including wound healing, vascular diseases (such as atherosclerosis, restenosis, and platelet activation), and inflammation and cancer. However, pharmacological regimens are often conceptualized primarily in terms of agonism or antagonism. A deeper mechanistic understanding of how GPCRs regulate selectivity, efficacy, and bias through allosteric pathways could therefore open new opportunities for next-generation pharmacology.
Previous structural and biochemical work has identified important microswitch motifs and a conserved sodium-binding pocket in class A GPCRs, but how these local elements connect to global activation and coupling across the receptor-G protein complex has remained unclear. To address this problem, the present study integrates Rigidity Transmission Allostery (RTA)-invented and developed by Dr. Adnan Sljoka (RIKEN)-with 19F-NMR and molecular simulations to directly track communication pathways across A2AR and into the receptor-G protein complex. The 19F-NMR experiments were led by Prof. R. Scott Prosser (University of Toronto), and the molecular simulations were performed by Dr. Duy Phuoc Tran and Prof. Akio Kitao (Institute of Science Tokyo) together with Dr. Andrejs Tucs (RIKEN).
Overview of Research Achievement
1. Integrating mathematical allostery modeling, 19F-NMR, and simulations
The study began with RTA, a mathematical framework based on rigidity theory (led by Dr. Adnan Sljoka) for probing allostery in protein structures. RTA predicted prominent allosteric communication pathways across an agonist-bound A2AR-G protein complex and identified substantial involvement of tryptophan residues along these pathways.
The team then used 19F-NMR (led by Prof. Prosser, University of Toronto) to experimentally test and validate these predictions by monitoring receptor conformational changes at two complementary levels:
- A TM6 reporter, which reports on global activation-state changes, and
- Fluorotryptophan reporters, which reveal local conformational changes at multiple tryptophan sites distributed across A2AR.
To connect experiment and mechanism, the team used molecular dynamics (MD) simulations (led by Kitao lab, Institute of Science Tokyo) and rigidity/geometry-based Monte Carlo simulations (performed by Dr. Andrejs Tucs, RIKEN) to characterize conformational ensembles and interpret how specific residues and microswitches contribute to long-range coupling.
The MD simulations showed how key microswitch regions and the sodium pocket reorganize across activation-related ensembles, providing a structural interpretation for the NMR-observed population shifts. The Monte Carlo simulations further supported this picture by efficiently sampling alternative receptor conformations, revealing plausible pathways and intermediate states that connect the inactive and activation-like ensembles.
2. Discovery of a tryptophan-rich allosteric network
A central finding is the identification of a tryptophan-rich network embedded within major allosteric communication pathways predicted by rigidity theory, linking the receptor's ligand-binding pocket to intracellular signaling regions and, ultimately, to the G protein nucleotide-binding machinery. This result suggests that tryptophans are not merely passive structural elements or convenient NMR probes; rather, they are functionally positioned to participate in the transmission of allosteric signals during GPCR activation.
The study highlights the importance of key tryptophan residues including the conserved toggle-switch residue W246 and an extracellular loop residue W143. Mutation experiments (W246Y and W143Y), together with 19F-NMR and simulations, showed that perturbations at these sites produce long-range effects on receptor conformational behavior, supporting their role in regulating allosteric communication.
3. Sodium egress promotes activation-related states
Another major outcome is a mechanistic explanation for the functional importance of the conserved sodium pocket in class A GPCRs. The researchers found that higher sodium concentrations stabilize the inactive receptor state, whereas lower sodium concentrations strongly promote activation-related conformational ensembles. Notably, sodium depletion strongly enhances a precoupled state, an intermediate receptor state thought to precede productive G protein coupling.
These results support the idea that sodium release from the conserved pocket is a key enabling step in activation, facilitating access to conformations required for signaling.
4. Broader significance
Because the sodium pocket and related activation motifs are broadly conserved across class A GPCRs, the findings are expected to be relevant beyond A2AR and may inform activation mechanisms in many other medically important receptors. More broadly, the study provides an integrated framework linking:
- ion-dependent regulation,
- microswitch dynamics,
- tryptophan-mediated allosteric communication, and
- receptor-G protein signaling.
Future Development
This work establishes a powerful integrative framework combining 19F-NMR, rigidity-theory allostery modeling, and simulation-based structural analysis to study GPCR activation and coupling mechanisms. In addition, the workflow provides a scalable route to generate experimentally validated data of allosteric pathways, dynamics and functional-state regulation-scarce but critical resources for training and benchmarking next-generation AI models. Expanding these datasets across GPCR families, ligands, ions, and disease mutants can help de-risk AI approaches that aim to predict receptor dynamics, coupling, and drug response beyond static structure. Ultimately, such AI-ready mechanistic datasets may accelerate state-selective and allosteric GPCR therapeutic design.
In the future, the same approach can be extended to:
- other class A GPCRs and additional GPCR families,
- different receptor-G protein pairings,
- disease-associated receptor mutants, and
- ligand-specific activation pathways, including biased signaling.
By identifying receptor-specific allosteric pathways and activation intermediates, this framework may help guide the rational design of allosteric modulators and next-generation GPCR therapeutics with improved selectivity and reduced side effects. More broadly, the research highlights how combining experimental biophysics with mathematical modeling and advanced simulations can address complex, dynamic problems in structural biology and drug discovery.
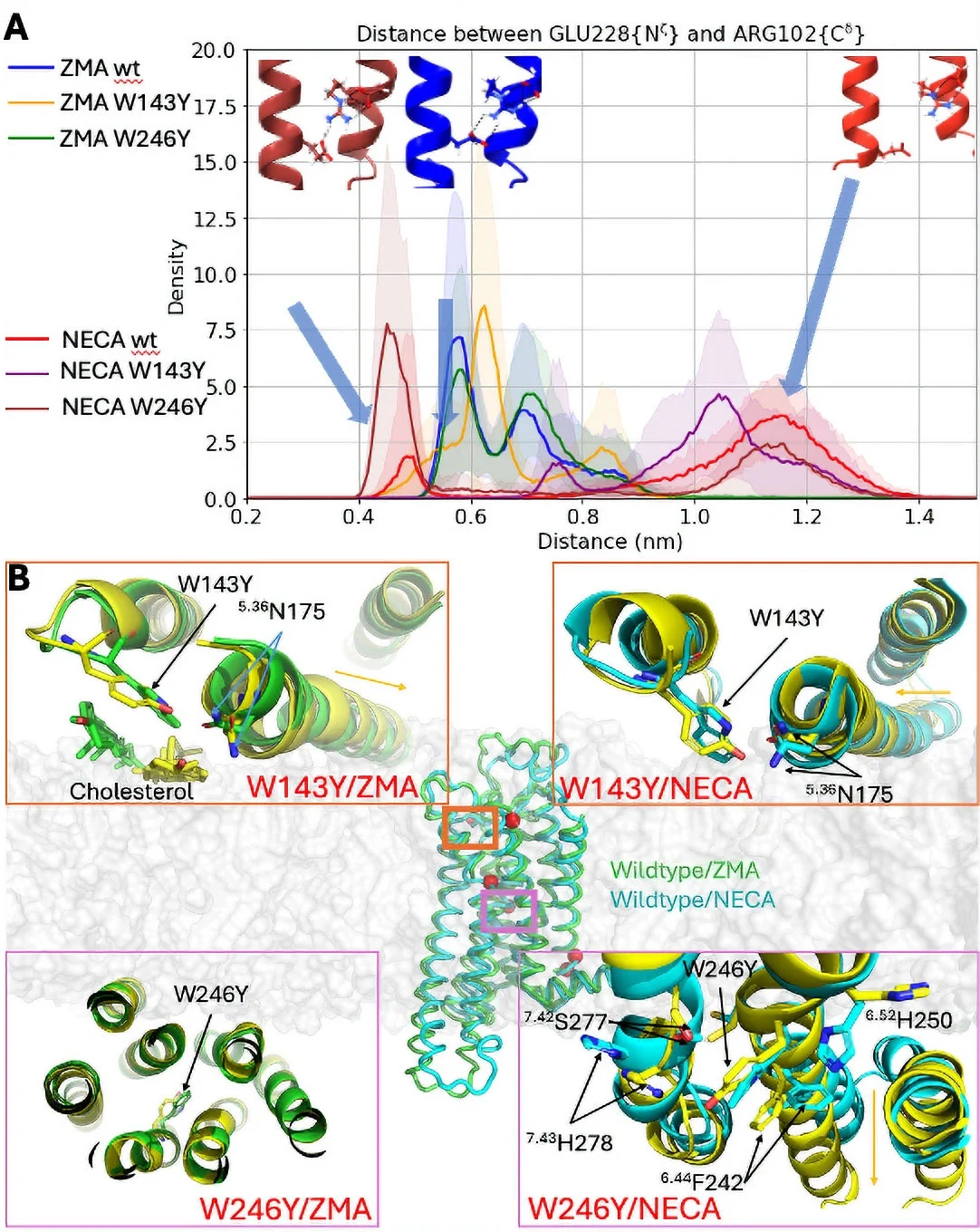
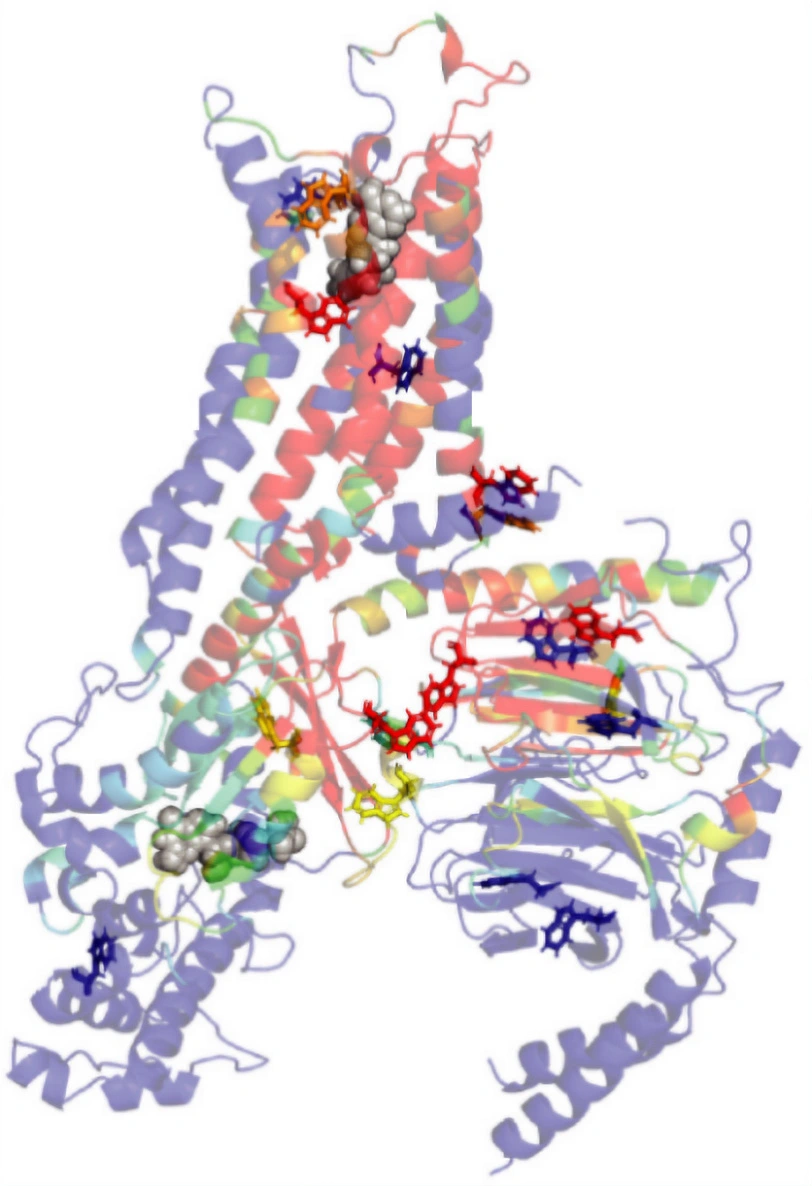
A. Distribution of the distance representing the conformational transition of a molecular micro-switch (on/off states). B. Representative structures of A2AR in complex with the agonist (NECA) and the antagonist (ZMA).
Reference
- Authors:
- Zhenzhou Qi1, Duy Phuoc Tran1, Andrejs Tucs1, Roopan Thiara , Louis-Philippe Picard , Koji Tsuda , Aditya Pandey , Adnan Sljoka* , Akio Kitao* , Robert Scott Prosser*
*corresponding authors
1Z.Q., D.P.T., and A.T. contributed equally to this work.
- Title:
- The Role of the Tryptophan-Rich Allosteric Network and Sodium Egress in GPCR Activation
- Journal:
- Proceedings of National Academy of Science, U.S.A.
Latest Article Publication Date: 2 March 2026






