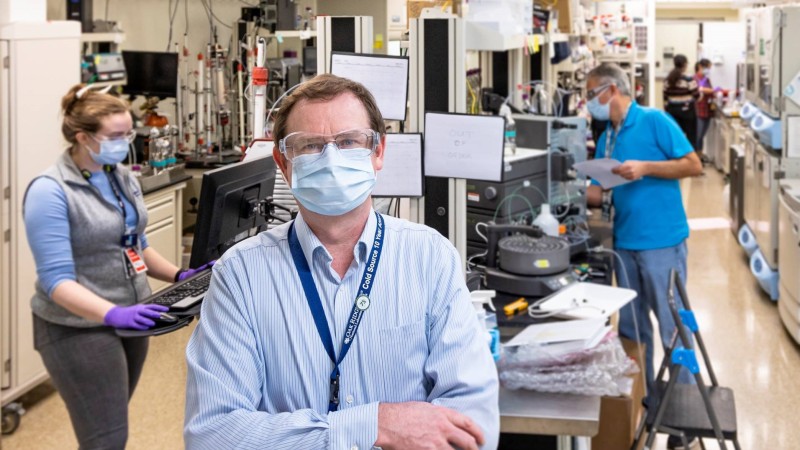

As the world grapples with the devastating effects of COVID-19, US national laboratories are mobilizing to use their facilities and resources to fight the deadly virus that causes the disease.
At the Department of Energy's (DOE's) Oak Ridge National Laboratory (ORNL), it's all-hands-on-deck for the world-leading experts in neutron scattering. Researchers at the lab's Spallation Neutron Source (SNS) and High Flux Isotope Reactor (HFIR) have a plan of attack to unleash a full barrage of neutron capabilities in an ambitious set of experiments that aim to provide critical pieces of information about the virus's biological structure and how it behaves. The neutron data will be combined with contributions from other national labs and sent to Summit-the world's most powerful supercomputer-to run simulations with the goal of finding inhibitors, or molecular matching compounds to block the virus's ability to replicate and spread.
"The coronavirus has been with us for many years in various forms and mutations. Today, it's a global pandemic and it's going to keep coming back again and again until we find a way to stop it," said Paul Langan, ORNL's associate laboratory director for Neutron Sciences. "So, what can we do? We can conduct research that provides us with insights that allow us to develop therapies and diagnostics that will make a lasting impact."
COVID-19 is the disease that comes from exposure to the virus SARS-CoV-2. It's part of the family of coronaviruses that have also led to diseases such as SARS and MERS, which cause acute respiratory and gastrointestinal or neurological diseases and other illnesses by infecting host cells and suppressing their immune responses. The surface of the virus is covered with spike-like tendrils that aggressively anchor it to cells in the body. Once it is attached, the genetic material encapsulated inside the virus hijacks the machinery of the host cell and forces it to produce viral proteins that lead to the propagation and spread of new viruses, which continue the assault by assimilating other cells.
"Neutrons are nondestructive particles that are highly sensitive to light elements such as hydrogen. When they interact with or bounce off atoms within a material, they reveal fundamental information about how the material's atoms are arranged and how the material functions," said Neutron Scattering Division Director Hans Christen. "That gives us the information we need to understand the function of such viral proteins at the level of individual atoms, but also to understand how such materials aggregate and interact with each other."
For the past several weeks, an ORNL team of molecular and structural biologists led by Hugh O'Neill, Director of the Center for Structural Molecular Biology (CSMB), has been developing a plan to study SARS-CoV-2 proteins that will be produced from synthetic DNA constructs. The genes will be inserted into bacteria to produce proteins of the virus, which will be studied using a suite of neutron scattering instruments to gain a better understanding of the structure and function of the disease. The new knowledge will lend itself to the development of improved methods for mitigating the virus.
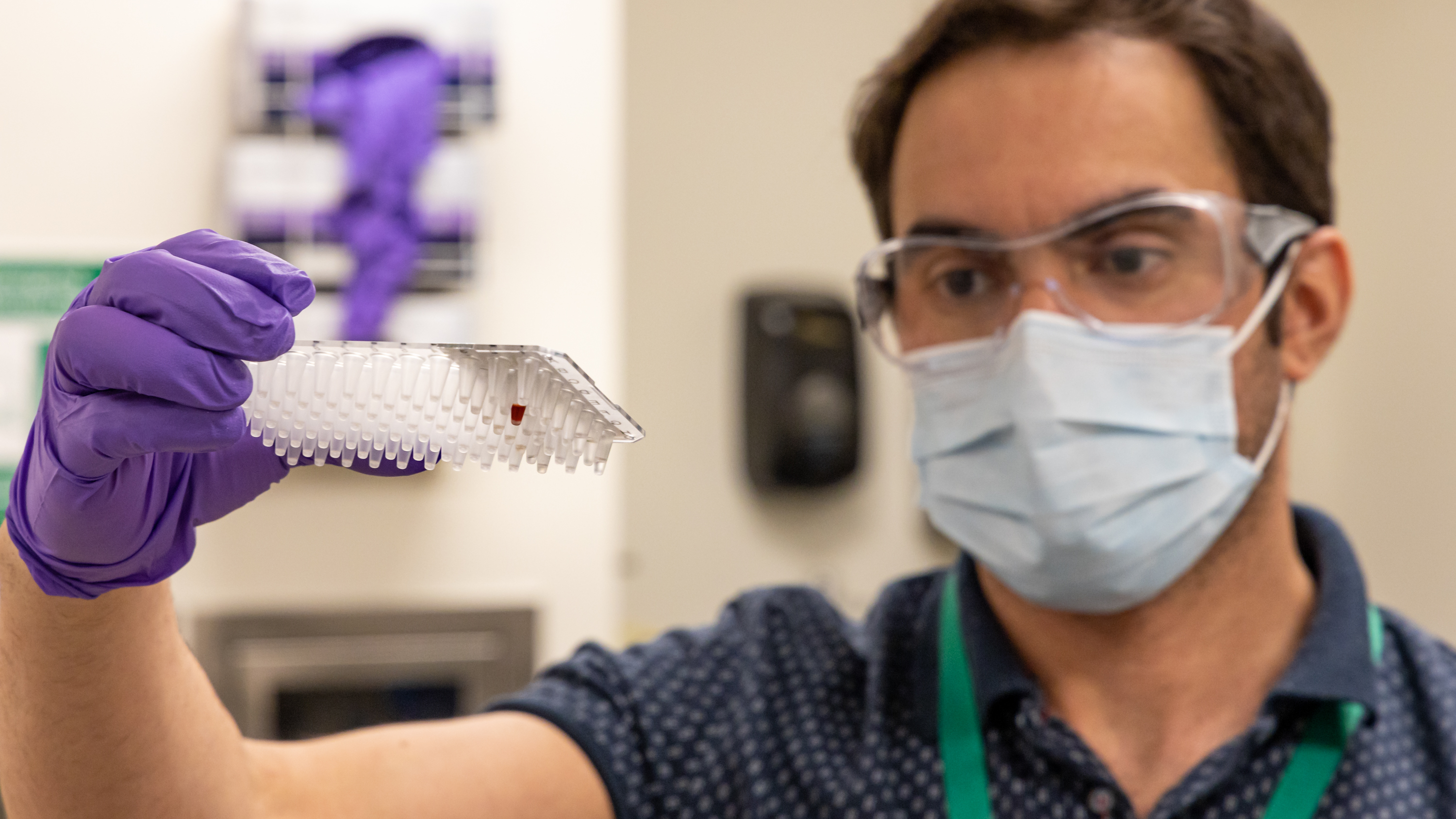

Postdoctoral researcher Wellington Leite is using small-angle x-ray scattering to characterize small components of SARS-CoV-2 including non-structural proteins NSP-7, NSP-8,
and NSP-15, the proteins that enable the virus to reproduce. Leite is part of a team of ORNL researchers at the Center for Structural and Molecular Biology that are synthesizing
these and other proteins associated with COVID-19 and preparing them for neutron scattering studies at ORNL's Spallation Neutron Source and the High Flux Isotope Reactor.
(credit: ORNL/Carlos Jones)
The team is also working in collaboration with other groups at ORNL and other institutions as well as DOE national labs that operate photon, electron, and computing facilities. The combination of those resources and techniques provide complementary data for improved drug therapies and for building more accurate computational models. Collaborators also include researchers from academia and the National Institutes of Health.
Data from the national laboratories is being published in open source journals as quickly as it becomes available to fast-track potential treatments.
Requesting rapid response access
Although initial beamline research will be conducted only by ORNL researchers, staff in the Neutron Sciences Directorate have developed a platform for external researchers to use to submit proposals with the expectation of a 2 day turnaround for approvals. Researchers requesting beamline access for mail-in experiments can submit proposals through the Rapid Access for COVID-19 Research portal. Limiting beamlines to internal staff is part of ORNL's efforts to keep employees safe while research is under way.
To be sure, there are no plans to bring the virus onto the Oak Ridge site. Only nonharmful components of COVID-19 are being synthesized and investigated, which pose no danger to the researchers and staff.
The task at hand
The ORNL neutron instruments will initially be used to study three components of the novel coronavirus: (1) the spike protein on the virus's surface, or the membrane shell; (2) the main protease responsible for forming the functional proteins that replicate the virus; and (3) the nonstructural proteins involved in the RNA replication process. These proteins are all from SARS CoV-2, the virus responsible for the COVID-19 disease. In addition, neutron scattering experiments will be conducted on small molecules associated with potential therapies currently being considered for the virus.
Both DOE neutron scattering flagship facilities are coming off planned maintenance cycles. After a successful outage, power was restored to the SNS accelerator on April 7, and the facility is expected to resume neutron production to begin COVID-19 research the week of April 13. HFIR is on track to restart April 28.
At present, ten SNS and HFIR instruments have been tapped to potentially conduct research into COVID-19: BASIS, Bio-SANS, CNCS, EQ-SANS, GP-SANS, IMAGINE, LIQREF, MaNDi, SEQUOIA, and VISION. The instruments represent four major neutron scattering techniques-diffraction, reflectometry, small-angle neutron scattering (SANS), and spectroscopy. Each technique will provide unique pieces of the complex puzzle at the atomic and molecular scales.
Diffraction
Neutron diffraction is used to provide the locations of individual atoms, which collectively provide an atomic portrait of a material's structure, as well as information about other formations and processes within the material.
The MaNDi and IMAGINE diffractometers will be used to study the main protease (SARS-CoV-2 protease), an enzyme responsible for cutting proteins that promote viral reproduction. Tracking the molecules' hydrogen atoms inside samples of crystallized proteins will provide insight into how this enzyme functions and how it binds to other small molecules, such as drug candidates.
"Because this protease protein is essential for the life cycle of the virus, that makes it a very attractive drug target. If we can design or retrofit current drugs to inhibit this, we can stop the virus from reproducing," said postdoctoral researcher Daniel Kneller, who joined neutron crystallographer Andrey Kovalevsky in January to do HIV research.
Small-angle neutron scattering
SANS is an ideal technique for studying large-scale structures, such as proteins or particles in solution, with sizes between 1 and 100 nanometers. SANS provides useful information such as the sizes and shape of molecules, as well as their various assemblies.
"The real power of neutron scattering lies in the fact that different biological molecules relevant to COVID-19-such as the proteins, RNA, and lipid molecules-have intrinsically different interactions with neutrons; and that allows us to distinguish among them when we make large-scale assemblies," said team lead and instrument scientist William Heller.
Using samples prepared in ORNL's CSMB laboratory, researchers are planning to use the three SANS instruments at ORNL to pinpoint and differentiate among molecular components such as RNA, phospholipids, and various hydrogenated and deuterated proteins and how they assemble to form the various complexes needed by the virus.
Reflectometry
"Similar to SANS, reflectometry looks at a molecule's gross properties, its size and shape and how it fits together, rather than looking at the atomic structure. The aspect that neutron reflectometry brings is that it allows us to study molecules in association with the membranes that they interact with," said instrument scientist John Ankner.
Research on the "old" coronavirus (SARS-CoV) revealed how the spike proteins expressed on the virus's surface, or lipid membrane, interact with and hijack healthy host cells, triggering membrane fusion. The Liquids Reflectometer can be used to observe structural changes in spike and other proteins as they interact with the host-cell membrane.
Lipid membranes form the boundary between cells and the outside world, explained Ankner. Membrane interactions of the virus and host cells can be measured by spreading the membranes across a thin water layer or silicon film. Drug molecules, or other chemicals, can then be introduced, whereupon neutrons will measure the nanoscale rearrangements to determine how the proteins function. Those findings, in turn, will inform the design of drug candidates that block the virus's ability to reproduce.
Spectroscopy
Neutron spectroscopy doesn't so much tell researchers where atoms are per se, but rather tells them how the atoms are behaving. Commonly used to study magnetic behavior and real-time chemical reactions, spectroscopy can be applied to understanding the dynamics of small molecules as well, such as those used in drug candidates currently being considered as COVID-19 inhibitors.
"The systems being considered will all have spectroscopic fingerprints which can be used to understand how the molecules are vibrating," said instrument scientist Matt Stone. "Neutrons are good at detecting those vibrations, which tell us a lot about the energetic forces between the atoms, how they move, and how they bind to other materials."
Stone says the spectroscopy experiments were inspired by a list of drug compounds developed via computational simulations performed by a team of researchers led by Jeremy Smith at the University of Tennessee (UT). Stone and other instrument scientists are planning to use the BASIS, CNCS, SEQUOIA, and VISION spectrometers at SNS to investigate small molecules from Smith's list of compounds to physically test their ability to bind with other agents. Each instrument provides unique details, and together the data sets will be used to build a more complete picture of the dynamics of each compound and its viability as a drug candidate.
COVID-19 is the disease caused by the virus SARS-CoV-2. The virus is constructed with "spike" proteins on its outer membrane that anchors and fuses with host cells. Once
attached, the genetic material inside the virus hijacks the machinery of healthy cells, creating new proteins that enable the virus to reproduce. (credit: ORNL/Jill Hemman)
Calculating novel coronavirus
Neutron scattering at ORNL will be used to generate structural experimental data that will be shared with teams across the laboratory and at other institutions. In particular, data sets will be sent about 1 mile down the road from SNS and HFIR to the Oak Ridge Leadership Computing Facility, home of the Summit supercomputer.
"I know from firsthand experience how the national laboratories have unique and synergistic capabilities that can be used to understand complex scientific problems," said instrument scientist Leighton Coates. "At ORNL, we have many core strengths, but we certainly do have strong capabilities in neutron sciences and supercomputing."
In addition to conducting neutron scattering experiments, Coates is working with ORNL computing teams as well as UT's Smith to make the most of their data. The IBM AC922 Summit-the world's fastest supercomputer-will be used to run a series of simulations using neutron scattering data to better understand the dynamics of the deadly virus, with the aim of identifying drug inhibitors that jam the molecular lock-and-key function the virus requires to reproduce. The massive computing power of the 200-petaflop system will enable the team to do in days to a week what would take months on a desktop computer.
"We have some of the most powerful tools the world has to offer for scientific research," said O'Neill. "Because coronaviruses have been around for some time, we have knowledge about the steps they take in reproducing themselves, which offers us ideas or clues on how we can combat the virus, stop it in its tracks, and save lives."






