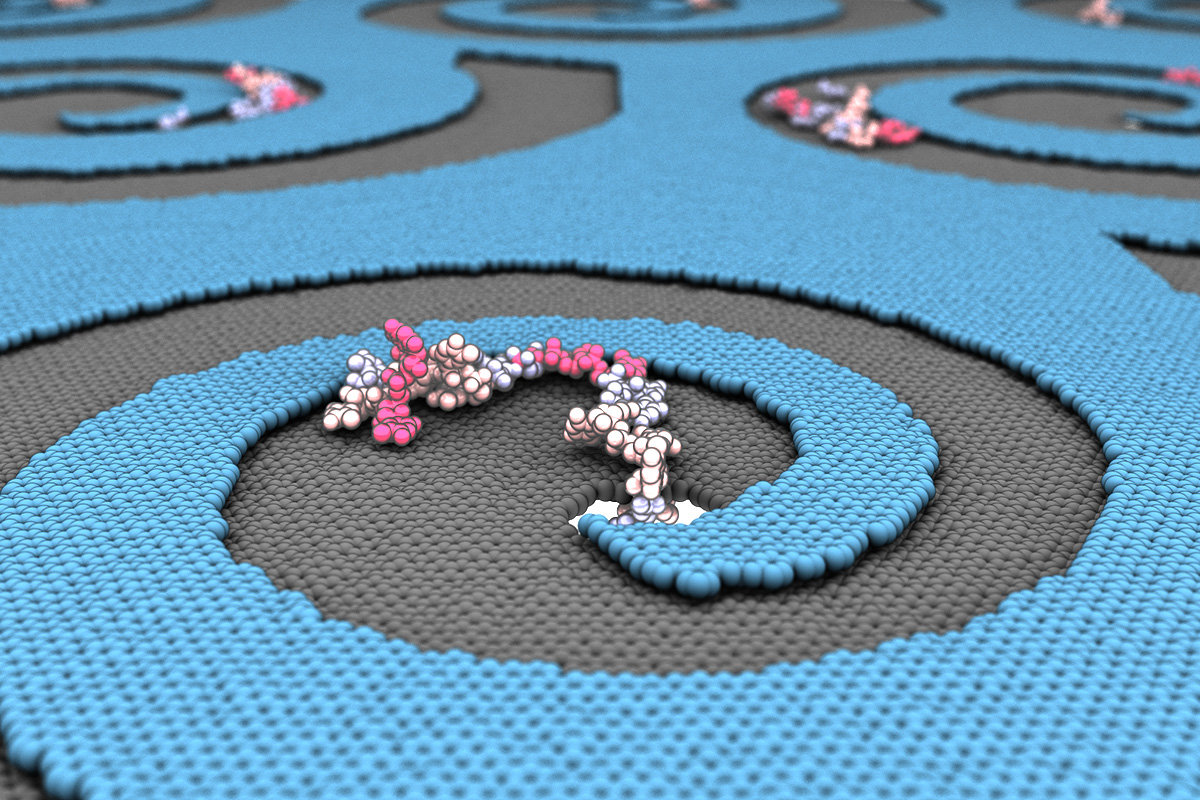

Laboratory-engineered membrane defects with edges that spiral downward would give biomolecules like DNA, RNA and proteins no other option than to sink into a nanopore for delivery, sorting and analysis.
Graphic courtesy Manish Shankla
CHAMPAIGN, Ill. — While watching the production of porous membranes used for DNA sorting and sequencing, University of Illinois researchers wondered how tiny steplike defects formed during fabrication could be used to improve molecule transport. They found that the defects – formed by overlapping layers of membrane – make a big difference in how molecules move along a membrane surface. Instead of trying to fix these flaws, the team set out to use them to help direct molecules into the membrane pores.
Their findings are published in the journal Nature Nanotechnology.
Nanopore membranes have generated interest in biomedical research because they help researchers investigate individual molecules – atom by atom – by pulling them through pores for physical and chemical characterization. This technology could ultimately lead to devices that can quickly sequence DNA, RNA or proteins for personalized medicine.

Physics professor Aleksei Aksimentiev, left, and graduate student Manish Shankla.
Photo by L. Brian Stauffer
In 2014, University of Illinois physics professor Aleksei Aksimentiev and graduate student Manish Shankla demonstrated a graphene membrane that controlled a molecule's movement through a nanopore by means of electrical charge. They discovered that once the molecules are on the surface of the membrane, it is very difficult to get them to shuffle into the membrane's pores because molecules like to stick to the surface.
While on sabbatical at Delft University of Technology in the Netherlands, Aksimentiev found that DNA tends to accumulate and stick along the edges of fabrication-formed defects that occur as linear steps spanning across the membrane's surface. The Illinois team's goal was to find a way to use these flaws to direct the stuck molecules into the nanopores, as a principle that can also apply to the delivery, sorting and analysis of biomolecules.
To refine and confirm their observations, the researchers used the Blue Waters supercomputer at the National Center for Supercomputing Applications at Illinois and the XSEDE supercomputer to model the system and molecule movement scenarios at the atomic level.

In this supercomputer simulation, a DNA molecule, directed by a force, moves down a fabrication-formed step defect along the surface of a graphene membrane.
Video courtesy Manish Shankla
"Molecular dynamics simulations let us watch what is happening while simultaneously measuring how much force is required to get the molecule to clear a step," Aksimentiev said. "We were surprised to find that it takes less force to move a molecule down a step than up. Although it may seem intuitive that gravity would make stepping down easier, it is not the case here because gravity is negligible at the nanoscale, and the force required to move up or down should be the same."
See a video at https://youtu.be/kWEc_597F0g
Aksimentiev said team members originally thought they could use concentric defect patterns that form around the pores to force the molecules down, but their simulations showed the molecules congregating along the edges of the steps. That is when it dawned on them: A defect with edges that spiral into a pore, combined with an applied directional force, would give the molecule no other option than to go into the pore – kind of like a drain.
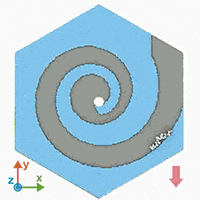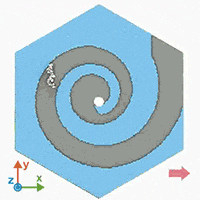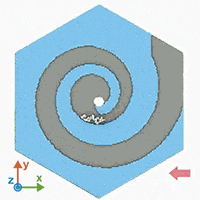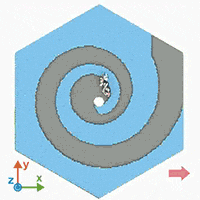
A supercomputer simulation shows a force-guided DNA molecule as it is directed down a spiral defect toward a graphene nanopore.
Video courtesy Manish Shankla.
"This way, we can drop molecules anywhere on the membrane covered with these spiral structures and then pull the molecules into a pore," he said.
See a video at https://youtu.be/VWlbbvl1ZSY
The researchers have not yet produced a membrane with spiral defects in the laboratory, but that task may be easier than trying to rid a graphene membrane of the current molecule-immobilizing step defects, they said.
"When manufactured at scale, defect-guided capture may potentially increase the DNA capture throughput by several orders of magnitude, compared with current technology," Shankla said.
"After a long development process, we are excited to see this principle used in a variety of other materials and applications such as delivery of individual molecules to reaction chambers for experiments," the researchers said.
The National Institutes of Health, National Science Foundation and the Dutch Research Council supported this research.






