Clethra fargessi Franch., in the family Clethraceae, is an important plant with good ornamental value and environmental adaptability that widely distributed in mountainous areas of Central China. So far, Clethra delavayi Franch. is the only complete chloroplast genome that has been sequenced with paucity in the available DNA fragments of Clethra. Therefore, there is a need to sequence more chloroplast genomes of Clethra species for further study for the family.
The East African Flora and Taxonomy Research Group from the Wuhan Botanical Garden reported the complete chloroplast genome of C. fargesii and comparative analysis for the genome features of Clethra species with the closely related species in Ericales.
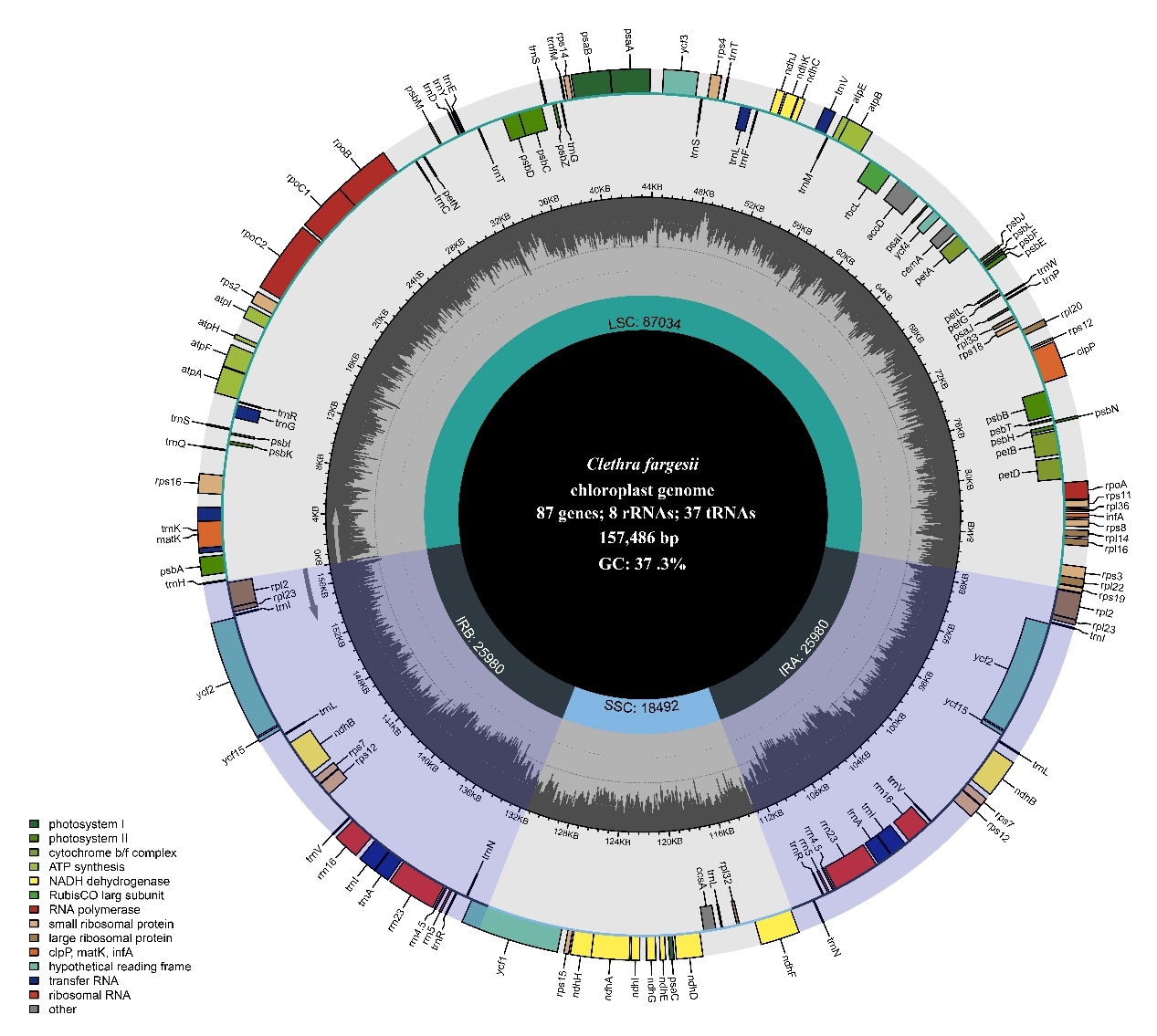
The complete chloroplast genome of C. fargesii is 157,486 bp in length, annotated with 112 single-copy genes, including 80 protein-coding genes, 30 tRNA genes, and four rRNA genes.
The number of genes, GC content and codon usage bias of C. fargesii, are highly similar to the other family species in the order Ericales. Also, the chloroplast genome is highly conserved in some aspects by alignment and rearrangement, selection pressure, and nucleotide diversity analysis.
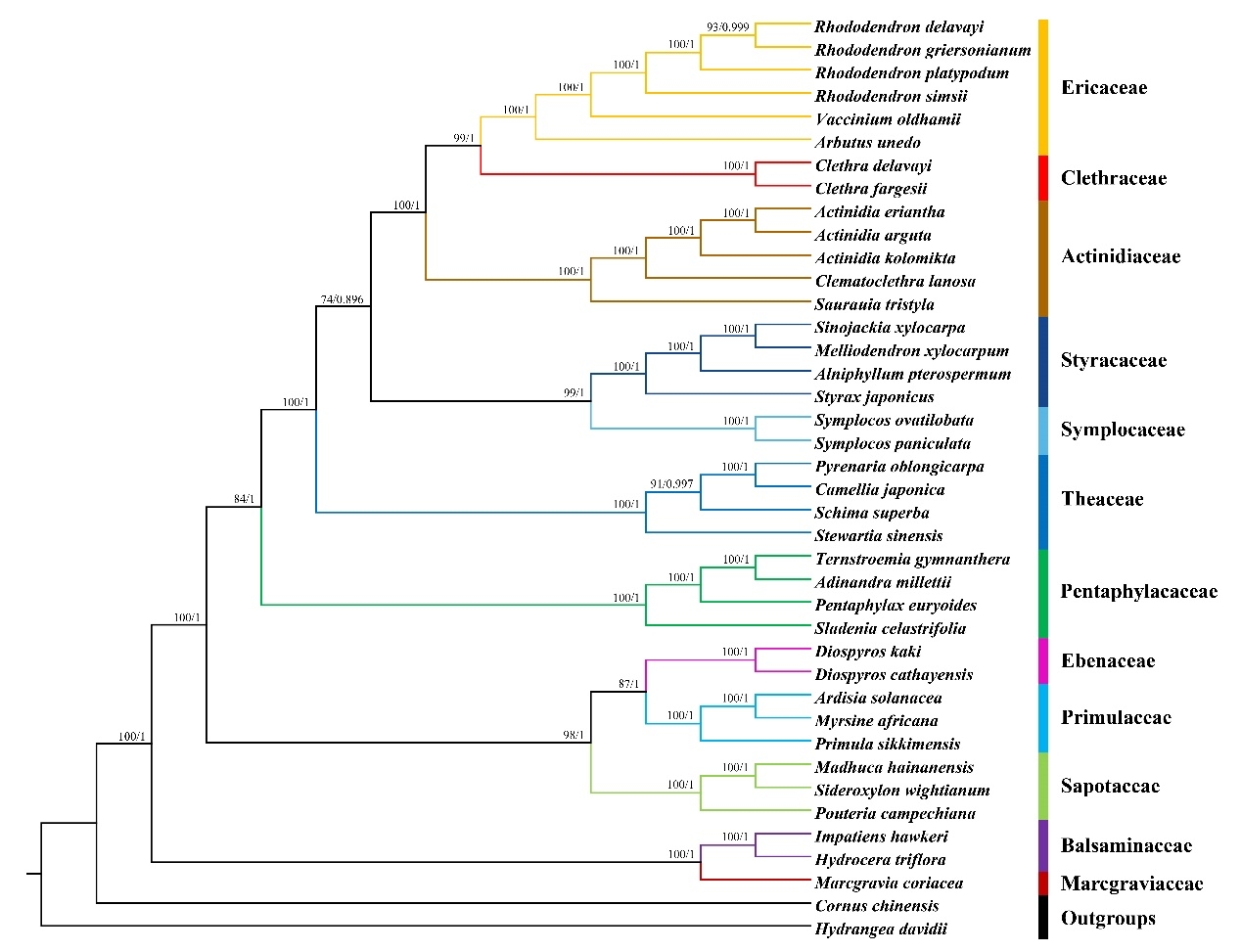
73 microsatellite sequences (SSR) and seven regions with high nucleotide diversity are identified in this genome, which could be used as potential markers for the identification and phylogeny of Clethra species. The phylogenetic tree in Ericales is reconstructed based on the 75 protein-coding sequences of the chloroplast genome, which supports Clethra and Ericaceae are the sister groups.
This study provides important data for the study of chloroplast genomes of Clethra and will be of great significance for the phylogenetic study and species evolution of this genus.
The study, entitled "Complete Chloroplast Genome of Clethra fargesii Franch., an Original Sympetalous Plant from Central China: Comparative Analysis, Adaptive Evolution, and Phylogenetic Relationships", was published in forests.