Alzheimer's disease (AD) is the most common type of dementia. It is a progressive disease beginning with mild memory loss and possibly leading to loss of the ability to carry on a conversation and respond to the environment. It involves parts of the brain that control thought, memory, and language and can seriously affect a person's ability to carry out daily activities. AD has become an increasingly severe public health and social problem. According to the Alzheimer's Association, one patient is diagnosed with AD every 3 seconds worldwide, and in China, there are an estimated 300,000 new cases every year.
The research team led by Prof. ZHOU Ruhong from the Zhejiang University Institute of Quantitative Biology recently published an article entitled "Molecular Mechanism of Secreted Amyloid-β Precursor Protein in Binding and Modulating GABABR1a" in the journal Chemical Science, offering insights into the future development of relevant therapeutics.
Amyloid-β peptide is the main component of senile plaques found in the brains of patients with AD and is derived from the normal proteolytic cleavage of the amyloid-β precursor protein (APP). A recent phenomenal study revealed that the extension domain of secreted amyloid-β precursor protein (sAPP) can bind to the intrinsically disordered sushi 1 domain of the γ-aminobutyric acid type B receptor subunit 1a (GABABR1a) and modulate its synaptic transmission. This work has laid an important structural foundation for the modulation of GABABR1a; however, the detailed molecular interaction mechanism, crucial for future drug design, remains elusive.
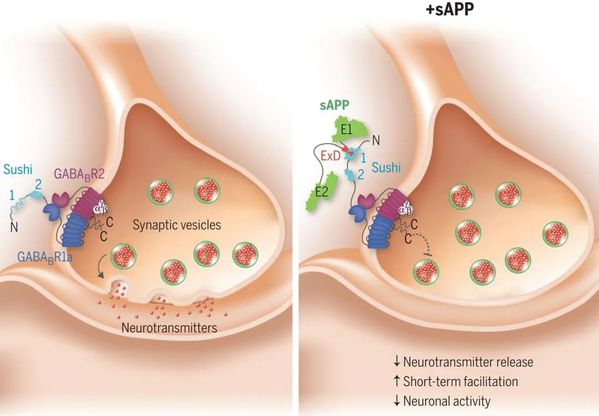
Structural comparison of sushi 1 domain bound to WT APP 9-mer (left) and mutated APP 9-mer's (right)
In this study, the researchers further investigated the dynamic interactions between sAPP peptides and the natively unstructured sushi 1 domain using all-atom molecular dynamics simulations, for both the 17-residue sAPP peptide (APP 17-mer) and its minimally active 9 residue segment (APP 9-mer). They explored the mutations of the APP 9-mer with rigorous free energy perturbation (FEP) calculations. Their in silico mutagenesis studies reveal key residues (D4, W6, and W7) responsible for the binding with the sushi 1 domain. More importantly, one double mutation based on different vertebrate APP sequences from evolution exhibits a stronger binding, indicating a potentially enhanced GABABR1a modulator.
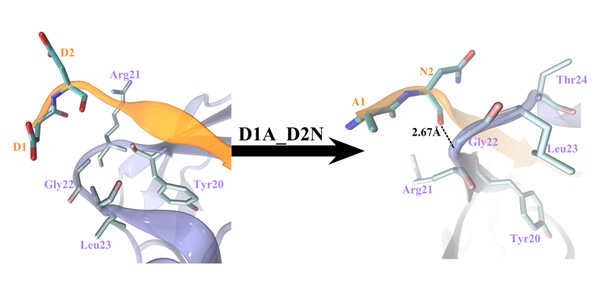
The binding complex structural changes due to the double mutation D1AD2N of APP 9-mer
"These large-scale simulations may provide new insights into the binding mechanism between sAPP and the sushi 1 domain, which could open up new avenues in the development of future GABABR1a-specific therapeutics," said Prof. ZHOU Ruhong.






