AP2/ ERF superfamily is one of the largest transcription factor families in plants and plays important roles in the diverse processes. In long evolution process, the functions and numbers of AP2/ ERF changed a lot. Flowering as a hot topic addressed numerous studies, which was also founded to be association with AP2/ ERF genes. However, little is known about the importance of AP2/ ERF genes for flowering in Actinidia eriantha (kiwifruit).
A research team at the Wuhan Botanical Garden of the Chinese Academy of Sciences identified and characterized AP2/ ERF family from the A. eriantha genome and explained the evolution of AP2/ ERF gene family in two main kiwifruit species. Several key regulators in flower development were predicted as well.
A total of 158 AP2/ ERF genes were identified and they were divided into four major subfamilies in A. eriantha. These genes demonstrated a favorable collinearity within A. eriantha, and many of AP2/ ERF genes experienced duplication events and were undergoing a purifying selection. Gene structure and protein motif analysis showed that AP2/ ERFs in different families were more conservative.
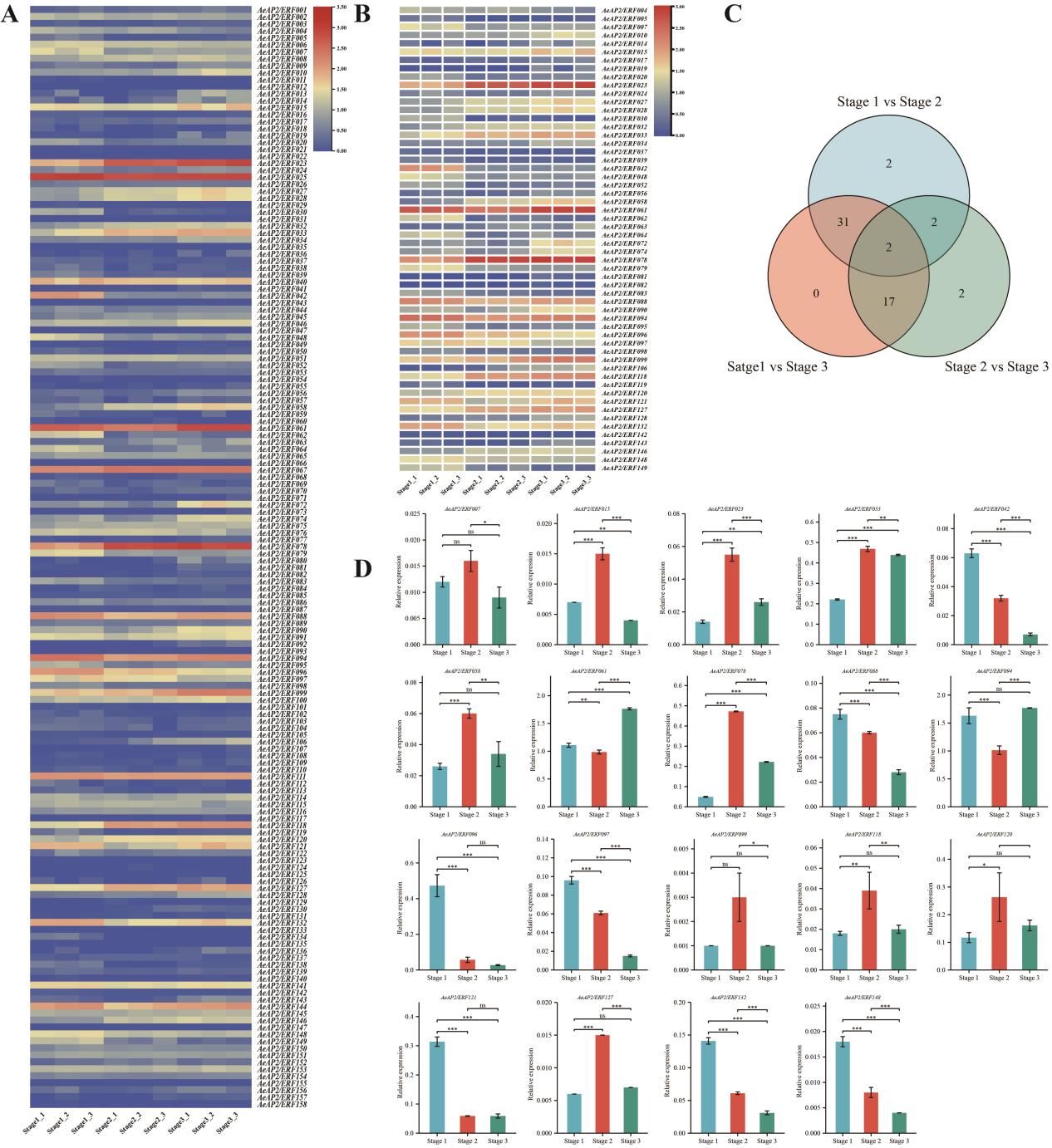
Besides, one third of AeAP2/ ERFs were strongly associated with flower transition by RNA- seq of different flower stages. Among them, two genes were expressed abundantly, which indicated that they may act a vital role in plant flower development regulation and flower tissue forming.
This study displays the first comprehensive analysis of AP2/ ERF genes in kiwifruit, and it would provide help for screening genes for further functional identification and for genetic improvement of agronomic traits of kiwifruit.
This work was funded by the National Natural Science Foundation of China and the results have been published on BMC Genomics entitled "Genome-wide identification and characterization of AP2/ ERF gene superfamily during flower development in Actinidia eriantha."