In their landmark 1961 paper of the lac operon, Nobel laureates François Jacob and Jacques Monod speculated that RNA might control gene activity in bacteria through base-pairing interactions. But once protein transcription factors were discovered, the idea was tossed aside.
Sixty years later, VP&S biologists now show that Jacob and Monod were on to something.
Some species of bacteria, Tanner Wiegand, Samuel Sternberg, and colleagues report in Nature, have evolved an RNA-guided gene activating system by transforming a copy of a CRISPR-Cas gene-cutting system.
"Instead of cutting at a point defined by the guide RNA, this system instead drags along the transcriptional machinery of the cell to turn on genes," says Sternberg, professor of biochemistry and molecular biophysics and investigator of the Howard Hughes Medical Institute.
"It is a completely new category of gene regulation that upends the dogma that gene activation is controlled by protein-DNA recognition."
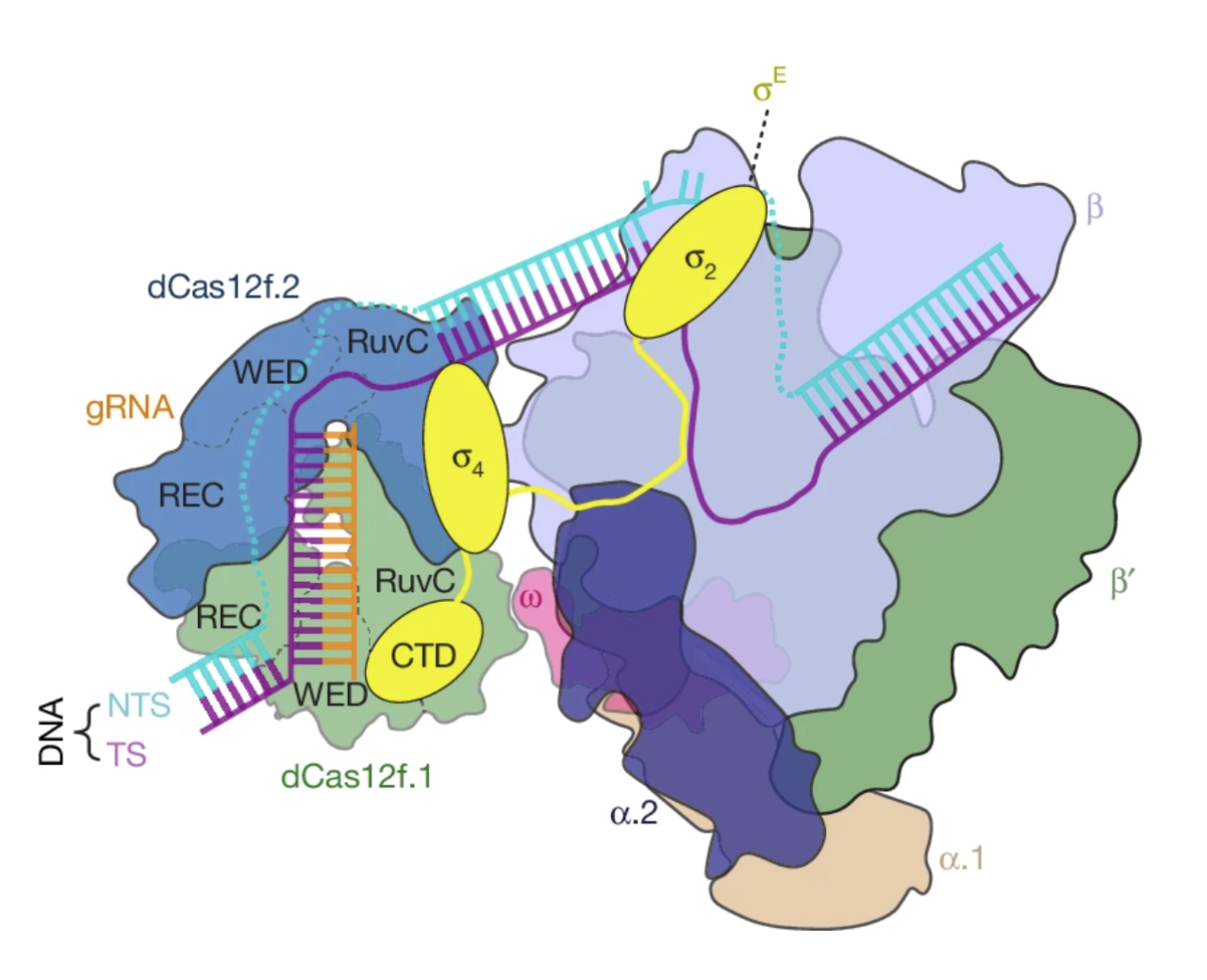
The newly discovered system, called dCas12f-σE, uses guide RNA (orange) to initiate transcription at a specific location in the bacterial genome and could be used to treat gastrointestinal diseases by activating therapeutically helpful genes in the gut microbiome. Image from Xiao, R., et al. (2026).
The system, called dCas12f-σE, was discovered in Bacteroidetes bacteria, common bugs in the human gut microbiome. dCas12f-σE turns on genes that are likely involved in the import of iron, peptides, and carbohydrates into the cell, and may provide the bacteria with more precise control over those systems. "It shows just how remarkable bacteria are at creating new uses out of older RNA-guided mechanisms, highlighting their untapped potential for genome control," adds Sternberg.
For synthetic biologists and genetic engineers, the system has wide-ranging biomedical applications and could be used in E. coli and other bacterial species to turn on genes in those genomes, a capability that is currently limited.
"One of this system's most exciting features is that it doesn't depend on the canonical motifs in DNA required by transcription factors," says Wiegand, a bioinformatics specialist in the Sternberg lab. "All we need to do is change out the guide RNA to reprogram this system to turn on any gene in the genome. It offers a lot more flexibility."
Sternberg's lab can now deploy multiple guide RNAs to turn on several genes at once, which may help in the search for new antibiotics.
"Certain bacteria are chock full of gene clusters that produce antimicrobial compounds, but we don't know how to turn them on," says Sternberg. "With an RNA-guided transcription system, we can turn on all the genes across the cluster and reawaken some of these cryptic pathways."
By activating therapeutically helpful genes in the gut microbiome, dCas12f-σE also has the potential to aid the treatment of gastrointestinal diseases
Though it may be difficult to reengineer the bacterial system to activate genes in mammalian cells, it's possible that similar systems exist in other eukaroyotes and could be tapped for new genome and transcriptome engineering applications.
"This superfamily of RNA-guided proteins is found pervasively in eukaryotes, in fungi, insects, and single-celled organisms. It's reasonable to envision that some species could have followed the same evolutionary arc of repurposing this very powerful molecular machine to turn genes on or off," says Sternberg.






