Human cells use large parts of their proteome for ribosome synthesis. In a manuscript published in NAR, the Kutay group exploited a genome-wide screening approach to identify factors required for 60S subunit biogenesis, which revealed an unexpected role of polyamine metabolism in 60S maturation.
The assembly of ribosomes is a complex process supported by hundreds of ribosome biogenesis factors (RBFs). Mutations in several RBF genes are associated with congenital diseases and tumorigenesis. While the cellular machinery supporting human 40S subunit synthesis has been well studied, far less is known about factors required for 60S synthesis in human cells.
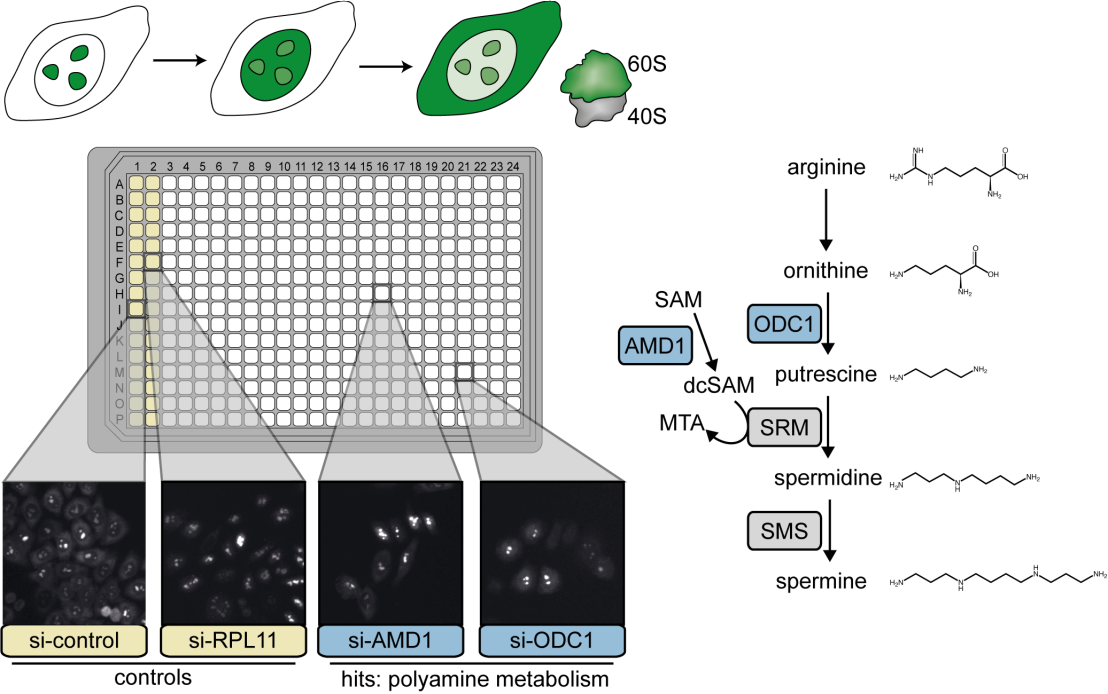
To identify such factors, the Kutay lab in collaboration with the Horvath (BRC, Szeged) and Zamboni group (IMSB, ETHZ), performed a genome-wide RNAi screen in human cells using a microscopy-based single cell readout. They identified 310 high-confidence factors required for 60S maturation, highlighting the conservation of the pathway from yeast to human and its interconnectivity with other key cellular processes. Also, several novel factors previously not implicated in ribosome maturation were identified, among them metabolic enzymes active in polyamine synthesis. Polyamines had been used as buffer additives in RNA and ribosome biochemistry for decades, yet links between polyamine metabolism and ribosome production in living cells had so far remained unexplored. Using a combination of cell biological and biochemical techniques, Dörner et al. show the importance of polyamine metabolism for 60S but not 40S subunit assembly, showcasing a novel function of polyamines. Their discovery leads to the exciting hypothesis that polyamines have a role in rRNA folding in vivo or even serve as structural components of 60S subunits.
Link to the paper in Nucleic Acids Research (NAR)call_made






