Genome-editing technologies using CRISPR-Cas nucleases have revolutionized plant science and hold immense promise in crop improvement. Conventional transgene-mediated CRISPR-Cas reagent delivery methods may be associated with unanticipated genome changes or damage, with prolonged breeding cycles involving alien DNA segregation and with regulatory restrictions regarding transgenesis.
DNA-free delivery has thus been developed by transfecting preassembled CRISPR-Cas9 ribonucleoproteins into protoplasts or in vitro fertilized zygotes. However, technical difficulties in regeneration from these wall-less cells make it infeasible to apply these approaches to most crop species. Alternatively, CRISPR-Cas ribonucleoproteins or RNA transcripts have been biolistically bombarded into immature embryo cells or calli to yield highly specific genome editing, albeit at low frequency.
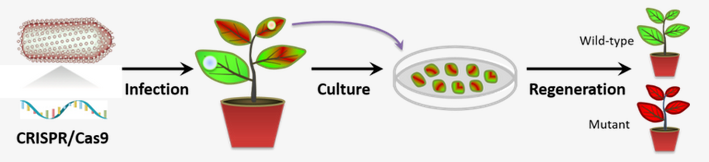
The research team headed by Prof. LI Zhenghe from the Zhejiang University College of Agriculture and Biotechnology reported the engineering of a plant negative-strand RNA virus-based vector for DNA-free in plant delivery of the entire CRISPR-Cas9 cassette to achieve single, multiplex mutagenesis and chromosome deletions at high frequency in a model allotetraploid tobacco host. Their research findings were published in the journal of Nature Plants on June 29.
In this study, over 90% of plants regenerated from virus-infected tissues without selection contained targeted mutations, among which up to 57% carried tetra-allelic, inheritable mutations. The viral vector remained stable even after mechanical transmission, and could readily be eliminated from mutated plants during regeneration or after seed setting.
This study provides a convenient, highly efficient and cost-effective avenue for CRISPR-Cas9 gene editing in plants through virus infection, thereby open the door to DNA-free plant genome editing.






