The research team led by Prof. CHEN Xuexin from Zhejiang University College of Agriculture and Biotechnology and the research team led by Prof. ZHAN Shuai from Chinese Academy of Sciences co-published a research article entitled "Two novel venom proteins underlie divergent parasitic strategies between a generalist and a specialist parasite" in Nature Communications, revealing the novel mechanism for diverse parasitic strategies among parasitoid species.
Parasitoid wasps have been extensively used as biological control agents for insect pests. They are highly diverse and exhibit substantial shifts in hosts or parasitic strategies even between closely related species. In response to the immune defense of the host, they have evolved two primary parasitic strategies: immune evasion and immune suppression. These two strategies can defeat host encapsulation and facilitate successful parasitism. However, the genetic bases underlying divergent parasitic strategies and how the host shift occurred has remained largely elusive.
Prof. CHEN Xuexin's research team has engaged in research into how parasitoid insects mediate host immunity, physiology, behavior and finally kill their host for nearly 30 years. In this study, they focused on one of the most representative systems of parasite-host coevolution-the genus Leptopilina (Figitidae), which is well known for parasitism of Drosophila-and discovered that two endoparasitoids, Leptopilina heterotoma (Lh) and L. boulardi (Lb) have developed different strategies to defeat the immune defense of Drosophila. Specifically, Lh adopts an active immune suppression strategy by destroying lamellocytes to inhibit host encapsulation and allow successful parasitism while Lb adopts a passive immune evasion strategy, whereby Lb eggs typically become attached to host internal tissues, which provide physical protection against complete encapsulation by host hemocytes. In addition, they also differ greatly in host range. Lb is specially adapted to D. melanogaster and its close relatives, whereas Lh can successfully parasitize a number of species across the Drosophila genus, causing these species to be labeled specialist and generalist parasitoid wasps, respectively. Thus, these two parasitic systems are an ideal model for research into the parasitic strategies of parasitoid insects and the adaptive mechanism in host range between parasitoids.
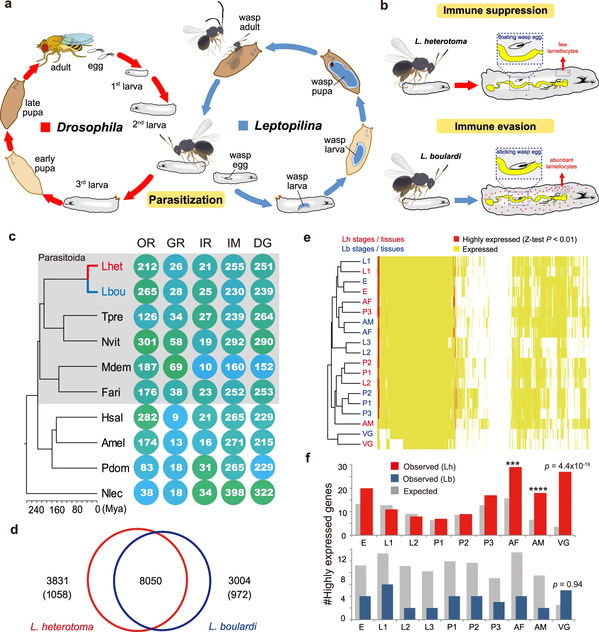
Researchers performed comprehensive studies in genomes, transcriptome, and proteome that enabled them to identify Lar and Warm as the most extremely outlier underlying the genetic differences between Lh and Lb. They verified their functions in vivo via RNAi experiments, which clearly showed that Lar and Warm are responsible for an active immune suppression strategy of Lh and a passive immune evasion strategy of Lb, respectively. To be specific, Lar triggers the lysis of the Drosophila lymph glands, eventually leading to the inhibition of host encapsulation response, whereas Warm contributes to the attachment of wasp eggs to the internal organs of the host, providing physical protection against the complete encapsulation that helps parasites escape from the host cellular immune response.
In addition, researchers also found that both Lar and Warm very likely originated from lateral gene transfer (LGT) and underwent subsequent gene duplication and expression specialization in the venom glands (VGs).
So far, this research has released the reference genomes of Lh and Lb and a complete repertoire of Lh and Lb venom proteins. "It is the first time that specific genes have been characterized associated with divergent parasitic strategies," said CHEN Xuexin. "We also uncovered the evolutionary scenarios of these genes, which add to our understanding of both the rapid speciation across parasitoids and the parasitoid-host coevolution."






