
The cryo-EM structure of the bacterial flagellar motor-hook complex (Cover image)
Many bacteria are capable of swimming 60 or even 100 times their own length in a second. This mobility constantly outdistances the cheetah-the fastest animal on the planet. Since Van Leeuwenhoek, known as the Father of Microbiology, first observed mobile bacteria in the 17th century, bacterial mobility and their mechanisms have attracted extensive research interests. A host of microbiologists, biochemists and biophysicists have committed their studies in this field, but many questions remain elusive.

Diagram of Salmonella Typhimurium and its flagella
Recently, TAN Jiaxing et al. from the Zhejiang University Life Sciences Institute and School of Medicine determined an atomic-resolution cryo-electron microscopy (cryo-EM) structure of the bacterial flagellar motor-hook complex in the assembled state from Salmonella Typhimurium. This study provides detailed molecular insights into the structure, assembly, and torque transmission mechanisms of the flagellar motor. It unravels the mystery of bacterial swimming and swarming mobility, and presents new strategies for antibiotics development. These findings were published in Cell on April 20.
The motility of bacteria is based on a special protein organelle-the flagellum. Many bacteria carry one or more flagella. The structure of the flagellum is made of three distinct parts: the motor, the hook and the filament. The flagellum is also an important weapon in bacterial pathogenesis.
The flagellar motor is one of the most complex protein machines and it can rotate at speeds of several hundred to over a thousand revolutions per second. Due to its complexity, the flagellar motor is a highly challenging topic in biochemistry and structural biology studies.
"To address this challenge, we aimed to solve its high-resolution structure. It is very difficult to purify the bacterial flagellar motor. It was too big!" said Dr. ZHU Yongqun, the project leader. To avoid the effects of the flagellar filaments on the purification of the motor, they genetically manipulated the genome of Salmonella Typhimurium. The mutant strain could normally produce the motor and the hook without the filament. After many attempts, the scientists designed a mild purification procedure for the motor particles and successfully obtained the intact flagellar motor-hook complex. They then collected the Cryo-EM images on an FEI Titan Krios electron microscope operated at 300 kV, and solved the atomic-resolution structure of the flagellar motor-hook complex in their collaboration with Dr. ZHANG Xing, Director of Cryo-EM Center of Zhejiang University.
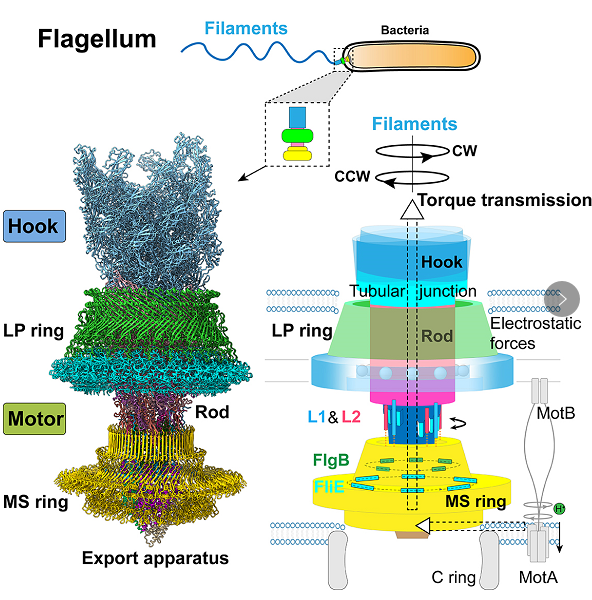
The structure of the flagellar motor-hook complex and its working model
How the flagellar motor is assembled and carries out the torque transmission for the fast rotation of the flagellum is a long-standing question. The structure reveals that the rod proteins in the motor make extensive intersubunit interactions via their distinctive structural elements, and form a compacted structure with high rigidity for the rod to mediate torque reception from the MS ring and torque transmission to the hook. Ten peptides protruding from the MS ring with the FlgB and FliE subunits of the rod mediate torque transmission from the MS ring to the rod and overcome the symmetry mismatch between the rotational and helical structures in the motor. The LP ring contacts the distal rod and applies electrostatic forces to support its rotation and torque transmission to the hook without significant structural obstacles and energy consumption.
The flagellum is one of the basic contents in the textbook of Microbiology. This study represents a landmark progress in the field. In the peer review process of the paper, a reviewer commented, "the cryo-EM structure represents a monumental structural effort". Another reviewer mentioned that "this work is outstanding and gives unparalleled resolution on some of the least understood parts of the flagellum". The study also indicates the diversity of torque transmission mechanisms of natural rotary protein machineries, and provides structural hints and information for the related researches on synthetic biology, biophysics, and nanomachines.

The research members of this study (From left to right: WANG Xiaofei, TAN jiaxing, ZHU Yongqun, XU Caihuang, ZHANG Xing, WU Hangjun, Wang Ting).






