New CRISPR tool eliminates undesired cells with ease. An international research team developed it. The Helmholtz Institute for RNA-based Infection Research was part of the team.
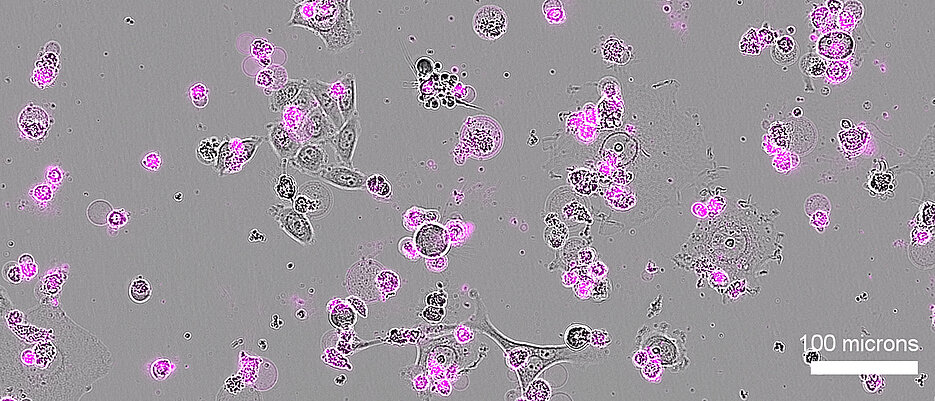
Many application - whether in medicine, biotechnology, or agriculture - require the ability to eliminate unwanted cells, since these can compromise health, reduce productivity, or interfere with desired biological processes. However, doing so without affecting other cells remains a significant challenge.
A collaboration of the Helmholtz Institute Würzburg (HIRI), Akribion Therapeutics in Zwingenberg, as well as the University of Utah and Utah State University in the US, has now resulted in a CRISPR-based tool that can target specific cells based on a recognized transcript, opening numerous potential uses. The findings were published in the journal Nature.
CRISPR technologies offer promising possibilities
A cell's identity and behavior are determined in part by which of its genes are currently being used and how active they are. Based on this activity pattern, cells can be distinguished and identified within heterogeneous populations. This is especially useful when certain cell types need to be removed, such as those associated with disease or those that have not undergone successful gene editing. Identifying these cells allows for targeted interventions but eliminating them often depends on the specific context and therefore requires customized approaches. This makes robust and versatile methods for detecting and selectively eliminating such cells highly sought-after tools in basic research, medicine, biotechnology, and agriculture.
When it comes to bacteria, so-called CRISPR technologies already offer promising possibilities for detecting and eliminating specific microbes. They are based on CRISPR-Cas, a bacterial immune system in which a ribonucleic acid (RNA) specifies the target sequence and a Cas protein (from "CRISPR-associated") acts as a nuclease - that is, an enzyme that cuts DNA. If the guide RNA is selected so that it matches only the DNA of a specific bacterium, that bacterium's genetic material is precisely recognized. The cut in the genetic material causes severe damage, and the affected bacterium dies.
A distinct strategy to eliminate specific cells
However, using the same nucleases in eukaryotes - organisms whose cells possess a nucleus containing DNA - has proven significantly more difficult. Researchers at HIRI, a site of the Braunschweig Helmholtz Centre for Infection Research (HZI) in cooperation with the Julius-Maximilians-Universität Würzburg (JMU), have teamed up with the company Akribion Therapeutics from Zwingenberg, as well as the University of Utah and Utah State University in the US, to apply these nucleases in eukaryotes - and subsequently found a distinct CRISPR strategy to eliminate specific cells.
Previous work involving the HIRI showed that the CRISPR nuclease Cas12a2 recognizes RNA target sequences, triggering non-specific cleavage of any nucleic acid it encounters-specifically RNA, single-stranded DNA, and double-stranded DNA. "This activity leads to extensive DNA damage in bacteria, causing a halt in growth and thus preventing the spread of a recognized invader," says Chase Beisel, affiliated department head at the HIRI. He is one of the corresponding authors of the new study. "In contrast to activated Cas9, which makes a single precise cut in the bound DNA, RNA target-activated Cas12a2 shreds all DNA it encounters, effectively killing the cell," says Ryan Jackson, Utah State University professor and co-corresponding author on the paper. "Its goal is not to correct anything. Instead, it's to destroy anything it sees," adds Yang Liu, further describing the nuclease Cas12a2. Liu is an assistant professor at the University of Utah and a corresponding author of the study.
However, it was unknown what would happen if Cas12a2 were triggered in eukaryotic cells - until now. The team of scientists and industry researchers found that Cas12a2 in yeast and human cells inactivated those containing the target transcript but spared those lacking the target sequence. "Cell death was sequence-specific, showed high sensitivity to mismatches, and occurred without any measurable unintended effects," says Beisel.
A wide range of applications
"Our technology provides us with a powerful tool for sequence-specific depletion of pathogenic cells," says Paul Scholz, co-founder and head of research and development at Akribion Therapeutics. Scholz is first and corresponding author of the publication. "In this study, we demonstrate the potential by targeting virus-infected cells as well as cancer cells transformed by a point mutation. However, our technology is programmable to target almost any RNA signature." Another possible application is to selectively remove unmodified cells to enrich successfully modified cells, thereby improving the quality of gene editing-something the team was also able to achieve in this study.
Given its in vitro activity - that is, under laboratory conditions and outside living organisms - the effect of activated Cas12a2 was to be expected. However, there was no guarantee of what would happen if it were released in a human cell: Initially, the team was concerned that Cas12a2 might eliminate cells other than the target cells if it were inadvertently triggered by RNA present elsewhere. However, this was not the case.
From oncology and chronic infections to gene editing: The ability to selectively eliminate cells based on their transcriptome opens new possibilities. "Because Cas12a2 can be programmed with a guide RNA to target any RNA sequence, and it shows little to no off targeting, we believe we have discovered a way to selectively kill cells across all of biology," Jackson says. "We show it can be used to enrich for gene editing, and to selectively kill cells harboring virus genes, and to kill cells with acquired mutations. We envision this technology will transform science, agriculture and medicine in ways previously unavailable."
Beisel adds: "We hope the research community will explore these new possibilities in greater detail, determining how they can be improved and applied beyond our study." The research team itself plans to continue developing Cas12a2 for clinical applications. At the same time, they will investigate ways to improve and expand the technology.
Funding
The study was supported by grants from the European Research Council, the R. Gaurth Hansen family, and the US National Institutes of Health.
Original publication
Scholz P, Thompson J, Crosby KT, Fauth T, Krah NM, Schlauderaff G, Back R, Berkheimer ZA, Jolley A, Sombroek D, Medert R, Zurek C, Dmytrenko O, Wilson E, Schut FT, Rutter J, Zhang X, Krohn M, Jackson RN, Beisel CL, Liu Y (2026). RNA-triggered cell killing with CRISPR-Cas12a2. Nature, https://doi.org/10.1038/s41586-026-10466-y






