Although autism is a common neurodevelopmental disorder, the multiple factors behind its onset are still not fully understood. Animal models of idiopathic autism*1, especially mice, are often used to help researchers understand the complicated mechanisms behind the disorder, with BTBR/J being the most commonly used mouse model in the world.
Now, an international research collaboration including Kobe University's Professor TAKUMI Toru and Researcher Chia-wen Lin et al. have made new discoveries regarding autism onset in mouse models.
In their detailed series of experiments and analyses of BTBR/J mice and the other subspecies BTBR/R, they revealed that endogenous retrovirus*2 activation increases a fetus's susceptibility to autism. They also discovered that BTBR/R exhibits autistic-like behaviors without reduced learning ability, making it a more accurate model of autism than the widely-used BTBR/J model.
It is hoped that further research will contribute towards better classification of autism types, as well as the creation of new treatment strategies for neurodevelopmental disorders*3.
These research results were published in Molecular Psychiatry on March 7, 2023.
Main points
- The researchers analyzed BTBR/J*4, a widely used mouse model of autism, and its subspecies BTBR/R*5 using MRI*6. This revealed that the corpus callosum*7, which connects the left and right hemispheres of the brain, was impaired in BTBR/J mice but not in BTBR/R mice.
- Genome and transcription analysis showed that BTBR mice have increased levels of endogenous retrovirus genes.
- Furthermore, single-cell RNA analysis*8 of BTBR/R mice revealed changes in the expression of various genes (including stress response genes) that are indicative of endogenous retrovirus activation.
- Even though BTBR/J and BTBR/R mice have the same ancestry, the results of various behavioral analysis experiments revealed differences in spatial learning ability and other behaviors between the two types of model mice.
Research Background
Autism (autism spectrum disorder) is a neurodevelopmental disorder that remains largely unexplored despite the rapidly increasing number of patients. Reasons for this continuing increase in people diagnosed with autism include changes to diagnostic criteria and older fathers becoming more common. Autism is strongly related to genetic factors and can be caused by abnormalities in DNA structure, such as copy number variations*9. Animal models, especially mice, are often used in research to illuminate the pathology of autism. Among these models, BTBR/J is a mouse model of the natural onset of autism that is commonly used. Studies have reported various abnormalities in BTBR/J mice including impairment of the corpus callosum (which connects the left and right hemispheres of the brain) and excessive immune system signalling. However, it is not fully understood why this particular lineage displays autistic-like behavioral abnormalities.
The aim of the current study was to shed light on the onset mechanism of these autistic-like behavioral abnormalities by conducting comparative analysis on BTBR/J and its subspecies BTBR/R.
Research Findings
First of all, the researchers conducted MRI scans on BTBR/J and BTBR/R mice to investigate structural differences in each region of the brain. The results revealed that there were differences between BTBR/J and BTBR/R mice in 33 regions including the amygdala. A particularly prominent difference discovered was that even though BTBR/J's corpus callosum is impaired, BTBR/R's is normal (Figure 1).
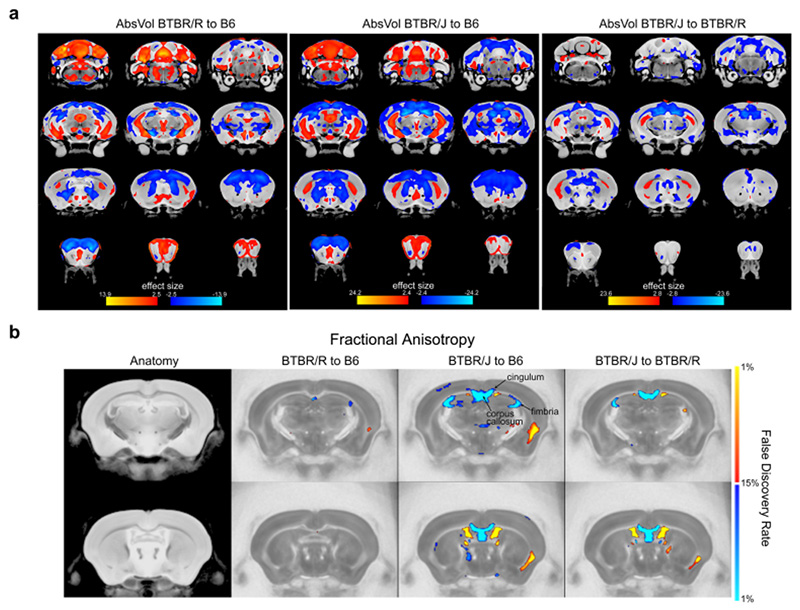
Figure 1: MRI scans showing brain structure differences in BTBR/J and BTBR/R mouse models of autism
a. Brain structure comparisons of the following mice: Left: BTBR/R and B6 (normal mouse), Center: Comparison of BTBR/J and B6, Right: BTBR/J and BTBR/R. b. Diffusion tensor imaging to compare differences in nerve fibers.
Red indicates the brain regions that were either bigger or had increased numbers of nerve fibers in BTBR/J mice in comparison to either B6 (left and center images) or BTBR/R (right image). Conversely, blue indicates brain regions in BTBR/J mice that were comparatively smaller or had decreased numbers of nerve fibers.
These scans revealed particularly significant differences between BTBR/J and BTBR/R mice's corpus callosum.
Next, the research group used the array CGH method*10 to compare BTBR/R's copy number variations with that of a normal mouse model (B6). They revealed that BTBR/R mice had significantly increased levels of endogenous retroviruses (ERV) in comparison to B6 mice (Figure 2 a-c). Furthermore, qRT-PCR tests revealed that these retroviruses were activated in BTBR/R mice (as shown in Figure 2d). On the other hand, in B6 mice there was no change in the expression of LINE ERV (which is classified in the same repetitive sequence), indicating that this retroviral activation is specific to BTBR (Figure 2e).
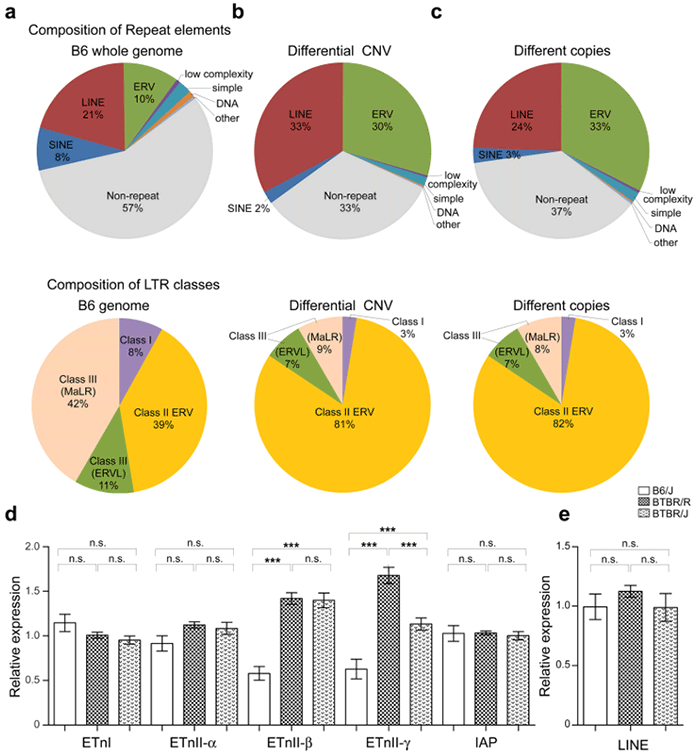
Figure 2: Copy number variation analysis and genetic expression analysis of BTBR/R autism model mice
a-c. Comparison of the repeated sequence of copy number variations, including retrovirus genes on the genome. d and e. Gene expression level analysis of repeated sequences: BTBR mice had more copy numbers of the endogenous retrovirus gene ERV than normal (B6) mice, and a portion of these genes were activated in BTBR mice.
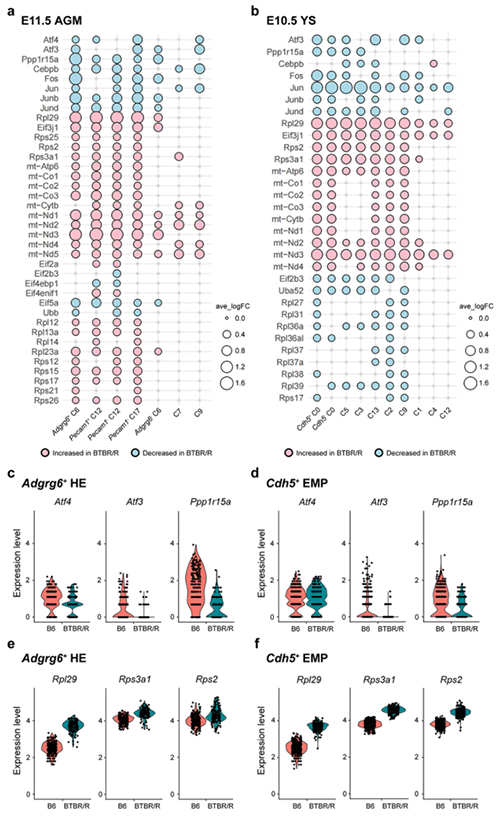
Figure 3: Single-cell RNA analysis of the yolk sac and the AGM region during embryonic development
a. AGM. B. Yolk sac: For BTBR/R autism model mice, genes with increased expression are indicated in pink and genes with decreased expression are in light blue.
c-f. Comparison of gene expression levels in each cluster.
In BTBR/R mice, there were changes in the expression of various genes (including stress response genes) that are indicative of endogenous retrovirus (ERV) activation.
Subsequently, the researchers carried out single-cell RNA analysis on the tissue of embryonic BTBR mice (on the AGM*11 and yolk sac*12). The results provide evidence of ERV activation in BTBR mice, as expression changes were observed in a group of genes downstream of ERV (Figure 3).
Lastly, the researchers comprehensively investigated the differences between BTBR/J and BTBR/R on a behavioral level. BTBR/R mice were less anxious than BTBR/J and showed qualitative changes in ultrasound vocalizations, which are measured as a way to assess communicative ability in mice (Figure 4a-f). BTBR/R mice also exhibited more self-grooming behaviors and buried more marbles in the marble burying test (Figure 4g, h). These two tests were designed to detect repetitive behavioral abnormalities in autistic individuals. From the results, it was clear that BTBR/R exhibits more repetitive behaviors (i.e. it is more symptomatic) than BTBR/J. The 3-chamber social interaction test*13, which measures how closely a mouse will approach another mouse, also revealed more pronounced social deficits in BTBR/R than BTBR/J mice (Figure 4i). In addition, a Barnes maze was used to conduct a spatial learning test, in which BTBR/J mice exhibited reduced learning ability compared to B6 (normal mice). BTBR/R mice, on the other hand, exhibited similar ability to B6 (Figure 4j).
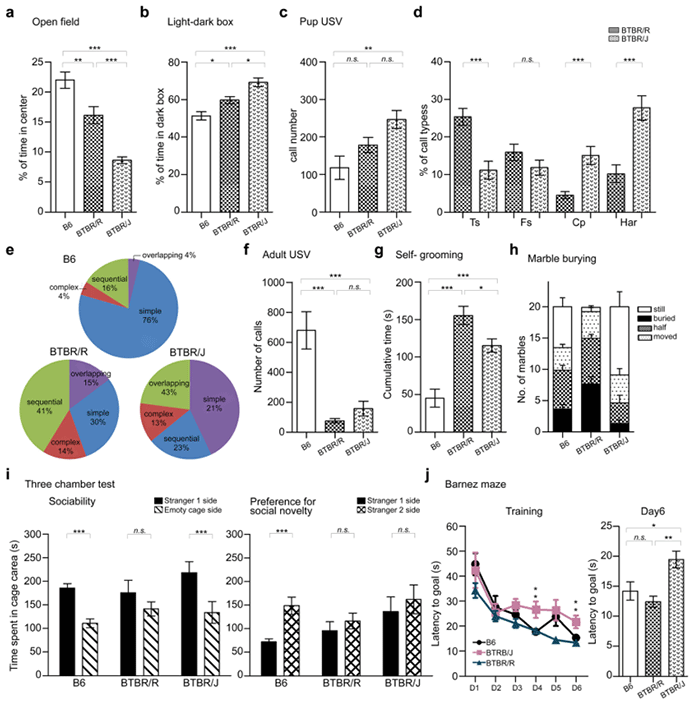
Figure 4: Detailed behavioral analysis of BTBR/J and BTBR/R autism model mice
a. Open field experiment b. Light-dark box experiment c-e. ultrasonic vocalizations made by mouse pups when separated from the mother. f. ultrasonic vocalizations of adult mice (in the presence of a mouse of the opposite sex). g. Self-grooming behavior. h. Marble burying test. i. 3-chamber social interaction test. j. Barnes maze spatial learning test.
BTBR/R and BTBR/J mice share key autistic-like behavioral abnormalities demonstrated in the results for experiments c through i. However, there are differences in anxious behavior (a, b) and spatial learning (j). BTBR/R mice did not demonstrate spatial learning difficulties. Therefore, BTBR/R is a more suitable model of autism than the existing BTBR/J model because it exhibited autistic-like behavior without compromised spatial learning ability.
Overall, the study revealed that retrovirus activation causes the copy number variants in BTBR mice to increase, which leads to the differences in behavior and brain structure seen in BTBR/J and BTBR/R mice (Figure 5).
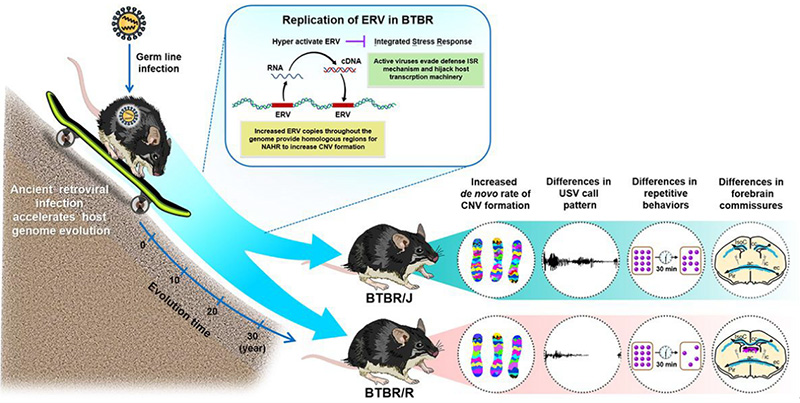
Figure 5: Diagram summarizing the findings of this study on differences in autism model mice
In BTBR autism model mice (BTBR/J and BTBR/R variants), retrovirus activation makes copy number variations occur easily. In other words, they could be said to evolve at a faster speed than normal mice. The supporting evidence for this is that although BTBR/J and BTBR/R mice share a common ancestry, a mere 30 years of being raised in different environments has led not only to behavioral differences between the two species but also significant differences in brain structure (i.e. BTBR/R mice have a functioning corpus callosum, while BTBR/J mice do not).
Further Developments
BTBR/J mice are widely used by researchers as a mouse model of autism. However, the results of this study highlight the usefulness of the other lineage of BTBR/R mice because they exhibit autistic-like behavior without compromised spatial learning ability. The results also suggest that it may be possible to develop new treatments for autism that suppress ERV activation. Furthermore, it is necessary to classify autism subtypes according to their onset mechanism, which is a vital first step towards opening up new avenues of treatment for autism.
Glossary
- 1. Idiopathic autism:
- Autism is considered to be a multifactorial disorder that can be caused by genetic and environmental factors. It is understood that genetic factors such as genetic and genomic abnormalities can cause autism, however, there are still many cases of autism where the cause is unknown. Autism where the cause cannot be specified (including environmental factors) is called idiopathic autism.
- 2. Endogenous retrovirus:
- A general term for an RNA virus with a reverse transcription. Most of these viruses are ancient and inactive and have been passed down through species over generations. About 8% of the human genome consists of these retroviruses.
- 3. Neurodevelopmental disorder:
- Previously called developmental disorder, this is a disorder that occurs in relation to a functional problem with the brain.
- 4. BTBR/J:
- A type of congenic mouse. Systemic behavior analyses of the BTBR line of mice have shown that it is the line that most closely resembles autistic behavior. Therefore, it is known as the idiopathic autism mouse model. This line is maintained and preserved by Jackson Laboratory (USA) and is widely used in autism research.
- 5. BTBR/R:
- This line of autism model mice has the same origin as BTBR/J. It was deposited at Riken Bioresource Research Center (Japan) in 1987 and has been maintained and preserved by the center ever since.
- 6. MRI:
- Magnetic Resonance Imaging. A non-invasive method that uses magnetic fields and radio waves to generate various cross-sectional images of the brain and other organs.
- 7. Corpus callosum:
- A bundle of commissural fibers. Commissural fibers connect the left and right hemispheres of the brain.
- 8. Single-cell RNA-seq:
- A method of comprehensively investigating the qualitative and quantitative aspects of all mRNA present in individual cells using a next generation sequencer. By combining this with statistical analysis methods such as dimension reduction, it is possible to classify cells based on their genetic expression and estimate the cell state. Furthermore, performing pseudo-temporal ordering analysis based on changes in the gene expression profile makes it possible to depict fibers in the cellular state that accompanies development.
- 9. Copy number variation:
- A copy number variation (polymorphism) is a phenomenon in which there is either less than one copy (deletion) or more than three copies (duplication) of genomic DNA that spans more than 1 kilobase (kb) on a chromosome. Normally there should be 2 copies. If there are 3 copies or just one copy, then this is a copy number variation. Copy number variations are found in 12% of the human genome. Copy number variations can exist in healthy, normal genomes but they can also cause genomic disease.
- 10. Array CGH method:
- This method can detect genome abnormalities including genomic DNA amplification and defects on a high resolution scale that encompasses all the chromosomes.
- 11. AGM:
- The Aorta-gonad-mesonephros (AGM) region is a hematopoietic site within the fetus (i.e. where cellular components of the fetus's blood are formed).
- 12. Yolk sac (YS):
- The yolk sac performs many critical functions that enable a fetus to develop, for example, it is involved in blood cell production (primary hematopoiesis).
- 13. 3-chamber social interaction test:
- An experiment designed to evaluate a mouse's sociability. Normally, mice are strongly interested in other mice. Therefore they spend more time in the chamber where there is another mouse than in the chamber that is empty.
Acknowledgements
The study was supported by funding from organizations including the following:
- Grants-in-Aid for Scientific Research (A) from the Japan Society for the Promotion of Science.
- The Japan Agency for Medical Research and Development's 'Strategic Research Program for Brain Sciences (SRPBS)' (Psychiatric & Neurological Disorders)
- Takeda Science Foundation
Journal Information
- Title
- "An old model with new insights: endogenous retroviruses drive the evolvement toward ASD susceptibility and hijack transcription machinery during development"
- DOI: 10.1038/s41380-023-01999-z
- Authors
- Chia-Wen Lin1,2,3, Jacob Ellegood4, Kota Tamada1,3, Ikuo Miura5, Mikiko Konda6, Kozue Takeshita6, Koji Atarashi6,7, Jason P Lerch4,8, Shigeharu Wakana5, Thomas J. McHugh2 and Toru Takumi1,3,9*
- 1 Laboratory for Mental Biology, RIKEN Brain Science Institute; Wako, Saitama 351-0198, Japan.
- 2 Laboratory for Circuit and Behavioral Physiology, RIKEN Center for Brain Science; Wako, Saitama 351-0198, Japan.
- 3 Department of Physiology and Cell Biology, Kobe University School of Medicine, Chuo, Kobe 650-0017, Japan.
- 4 Mouse Imaging Centre, Hospital for Sick Children, Toronto, Ontario M5T 3H7, Canada.
- 5 Technology and Development Team for Mouse Phenotype Analysis, Japan Mouse Clinic, RIKEN BioResource Research Center, Tsukuba, Ibaraki 305-0074, Japan
- 6 Department of Microbiology and Immunology, Keio University School of Medicine; Shinjuku, Tokyo 160-8582, Japan
- 7 RIKEN Center for Integrative Medical Sciences, Tsurumi, Yokohama 230-0045, Japan.
- 8 Wellcome Centre for Integrative Neuroimaging, University of Oxford, Oxford, Oxfordshire OX39DU, UK
- 9 RIKEN Center for Biosystems Dynamics Research, Chuo, Kobe 650-0047, Japan.
- * Corresponding author
- Journal
- Molecular Psychiatry






